COMUNICACIÓN
Phylogenetic analysis of Groundnut
ringspot virus isolates from peanut and
identification of potential thrips vectors in
peanut crop in Argentina
de Breuil, S.; F.R. La Rossa, A. Giudici, A. Wulff, N. Bejerman, F. Giolitti and S. Lenardon
S. de Breuil and N. Bejerman: Instituto de Patología Vegetal - Centro de Investigaciones Agropecuarias - Instituto Nacional de Tecnología Agropecuaria (IPAVE-CIAP-INTA), Camino 60 Cuadras Km 5 ½,
X5020ICA, Córdoba, Argentina; Consejo Nacional de Investigaciones
Científicas y Técnicas (CONICET). F. Giolitti and S. Lenardon: IPAVE-CIAP-INTA. F.R. La Rossa, A. Giudici and A. Wulff: Instituto de Micro-biología y Zoología Agrícola (IMYZA-CICVyA-INTA), Nicolás Repetto
y de los Reseros s/n, CP 1686, Hurlingham, Buenos Aires, Argentina.
Correspondence: debreuil.soledad@inta.gob.ar
SUMMARY
Groundnut ringspot virus (GRSV), genus Tospovirus, is a thrips-transmitted
virus infecting peanuts (Arachis hypogaea L.) in Córdoba province, Argentina.
Fourteen viral isolates were recovered from Tospovirus-like symptomatic plants
from different peanut fields. Viral isolates as GRSV were identified by serological
and molecular tests. Nucleotide and derived amino acid sequence analyses of
the nucleocapsid (N) gene indicated a high degree of identity between the
GRSV peanut isolates, indicating that there is no molecular variability in the N gene of the GRSV that infects peanuts in the cropping area of Córdoba. In this
study, we determined the presence of thrips species in the crop, which can
potentially transmit the virus. Thrips were observed in all the evaluated peanut
fields. Frankliniella schultzei was the most frequently identified species followed
by Caliothrips phaseoli and Frankliniella occidentalis. This work reports the
presence of F. schultzei and F. occidentalis in peanuts in Argentina for the first
time. These results along with the high degree of similarity between the GRSV
peanut isolates suggest that the virus could be transmitted by F. schultzei,
which has been cited as its most efficient vector.
Key words: Arachis hypogaea; Tospovirus; Molecular characterization;
Frankliniella schultzei
Análisis filogenético de aislamientos de
Groundnut ringspot virus desde maní e identificación de posibles trips
vectores asociados al cultivo de maní en la Argentina.
RESUMEN
Groundnut ringspot virus (GRSV, género Tospovirus) es un virus que
infecta naturalmente el cultivo de maní (Arachis hypogaea L.) en la región
productora de Córdoba, Argentina. En distintas localidades de la provincia, se colectaron 14 aislamientos virales provenientes de maníes que manifestaban
síntomas característicos de Tospovirus. Todos los aislamientos virales fueron
identificados como GRSV mediante pruebas serológicas y moleculares. El
análisis de las secuencias nucleotídicas y de amino ácidos deducidas del
gen de la nucleoproteína (N) reveló un alto grado de identidad entre los 14
aislamientos, indicando que no existe variabilidad molecular en el gen N del
GRSV que infecta maní en la provincia de Córdoba. En este estudio se determinó la presencia de trips en el cultivo que pueden potencialmente transmitir la
enfermedad. Estos insectos fueron observados colonizando maní en todos los
lotes evaluados. La especie identificada con mayor frecuencia fue Frankliniella
schultzei, seguida de Caliothrips phaseoli y Frankliniella occidentalis. Este es
el primer reporte de F. schultzei y F. occidentalis afectando maní en Argentina.
Estos resultados, junto con el elevado grado de similitud encontrado entre los
distintos aislamientos de GRSV, sugieren que el virus puede ser transmitido por F. schultzei, citado como el vector más eficiente del GRSV.
Palabras clave: Arachis hypogaea; Tospovirus; Caracterización molecular;
Frankliniella schultzei
Fecha de recepción: 07/10/14;
fecha de aceptación: 17/06/15
Argentina is one of the world's main peanut
(Arachis hypogaea L.) exporting countries.
The principal peanut growing area is located in
the center-south of Córdoba province (central
Argentina), accounting for approximately 91% of
the total production (Source: MAGyP - SIIA).
Peanut is affected by several tospoviruses
worldwide, including: Tomato spotted wilt virus
(TSWV), Groundnut ringspot virus (GRSV), Peanut
bud necrosis virus (PBNV), Peanut yellow spot virus
(PYSV) and the tentative species Peanut chlorotic
fan-spot virus (PCFSV) (Reddy et al., 1992; De
Avila et al., 1993; Satyanarayana et al., 1998; Chu
et al., 2001). TSWV, the type member, is the most
important species. It was first recorded in Texas
and has become one of the most serious peanut
diseases in the United States. To keep TSWV
incidence at low levels, it is necessary to combine
resistant cultivars with cultural measures such as
planting date, plant population, row pattern, tillage,
herbicide and insecticide application (Culbreath et
al., 2003). In the peanut growing area of Córdoba,
the crop is naturally affected by GRSV, an emerging Tospovirus whose infection causes plant stunting,
and smaller and distorted leaves showing severe
chlorosis and concentric ringspots (de Breuil et al., 2007; Pappu et al., 2009). GRSV was found to
be distributed in the central-northern area of the peanut-growing region of Córdoba, including the
departments of Río Segundo, Tercero Arriba, Gral.
San Martín and Juárez Celman (de Breuil et al.,
2008). The distribution is probably related to the
presence of insect vectors, since tospoviruses are
transmitted exclusively by thrips (Thysanoptera) in
nature (Whitfeld et al., 2005). GRSV is efficiently
transmitted in a propagative manner by the
thrips species Frankliniella schultzei (Tribom) and Frankliniella occidentalis (Pergande) (de Borbón & Gracia, 1996; Nagata et al., 2004) and by
Frankliniella gemina under experimental conditions
(de Borbón et al., 2006).
The aims of this work were to determine the
phylogenetic relationship between GRSV isolates
and to assess the presence of Tospovirus thrips
vectors in peanuts in the main producing area of
Argentina in order to generate epidemiological
information about GRSV disease.
The survey was carried out in the province of
Córdoba in February and March 2009 and 2010.
During the study, 92 farm fields were arbitrarily
selected and surveyed for virus infection. Young
leaf samples were collected from plants showing Tospovirus-like symptoms and stored at 4°C
until analyses. Samples were tested by doubleantibody sandwich-enzyme-linked immunosorbent assay (DAS-ELISA) using commercial ELISA kits
with specific GRSV/TCSV antibodies with their
respective positive and negative controls, following
the manufacturer's protocol (Agdia Inc., Elkhart, IN).
ELISA-positive samples were assayed by antigencapture (AC) followed by reverse transcriptionpolymerase chain reaction (RT-PCR). For AC,
PCR tubes were coated with 50 µl of a 1:200
(w/v) dilution of leaf tissue extract in carbonated
coating buffer, and incubated overnight at 4ºC.
RT-PCR was performed with two GRSV-specific
primers for the nucleoprotein (N) gene (Boari et al.,
2002) using the Access RT-PCR System (Promega
Corp., Madison, WI, USA). The following cycling
parameters were used: 48ºC for 45 min, 94ºC for
4 min, (30 cycles of 94ºC for 1 min, 50ºC for 2 min,
68ºC for 1 min), and a final extension of 68ºC for
7 min. Electrophoresis of the PCR products was
conducted on a 1.4% agarose gel and visualized
with 0.01% ethidium bromide under UV light. The
amplified fragments were purified using Wizard-PCR columns (Promega) and then ligated into
pGEM-T plasmid (Promega) according to the
manufacturer's instructions. Positive recombinant
clones were sequenced in both directions at
Macrogen Inc. (Korea Republic). The sequences
were compared with each other and with other Tospovirus sequences available in the GenBank
(NCBI, 2014). Multiple alignments of the nucleotide
and deduced amino acid sequences were obtained
using ClustalW and phylogenetic trees were
generated with the MegAlign program (DNASTAR,
Lasergene Software, 2001). Accordingly, the N
gene nucleotide and protein sequence analysis
have been selected as important criteria for Tospovirus species designation (King et al., 2012).
In addition, all the surveyed fields were visually
evaluated for presence or absence of thrips. Insect
sampling was made in 19 fields planted in different
locations and growing seasons. In each field,
a sample included 20 peanut flowers collected
randomly from the interior and margin of the plot,
or 20 leaves when flowers were not present. At
some sites, thrips were taken from weeds growing
within the crop. Samples were stored separately
in plastic bags; thrips were then transferred to
vials containing 70% ethanol using a brush, and
tagged with date and origin. All adult specimens
collected were individually mounted on slides and
examined under stereoscopic microscope with
60X magnifcation. Identifications were made by
confrontation with previously identifed material and
keys for Thysanoptera.
Fourteen Tospovirus-symptomatic peanut leaves
collected during 2009 from different locations across
the center-south of Córdoba province were positive to GRSV/TCSV infection according ELISA test.
During the 2010 cropping season, no Tospovirus-infected plants were detected. ELISA-positive
samples were assayed by RT-PCR and yielded an
expected 800-bp fragment corresponding to the N gene of GRSV. They were designated according
to their collection site as GRSV-ARo, GRSV-Baig,
GRSV-Beng, GRSV-Cab, GRSV-DVe, GRSV-Etr,
GRSV-Euf, GRSV-MGa, GRSV-Mol, GRSV-Oli,
GRSV-RCua, GRSV-RTer, GRSV-Tan and GRSV-TPu. Comparisons of nucleotide and deduced
amino acid sequences of the N fragment confirmed
their identities as GRSV. The 14 peanut isolates had
a high nucleotide identity to other GRSV sequences
(95.1-99.8%), showing the highest percentage
values with the Argentine GRSV isolate (GRSV-AR)
(Table 1). They also had identity values of 82.8-84.0% with TCSV and of 77.0-78.2% with TSWV.
Comparisons of amino acid sequences of peanut
isolates with other GRSV isolates revealed percent
identities ranging from 95.0% to 99.5%, whereas
with TCSV and TSWV, identities ranged from 86.9%
to 88.0% and 79.2% to 80.3%, respectively (Table
1). When the GRSV peanut isolate sequences
from Córdoba were compared with each other, the
percentages of identities ranged between 97.2-100% for nucleotides and 98.1-100% for derived
amino acid (Table 1), indicating a lack of molecular
variability in the N gene of the GRSV infecting
peanuts in the cropping area of Córdoba.
Table 1. Percent nucleotide identities (above diagonal) and derived amino acid identities (below diagonal) of the N gene of GRSV peanut
isolates with other reported tospoviruses.

The origin of the Córdoba peanut isolates of GRSV and the origin and the accession numbers of the tospovirus sequences available in
the GenBank are indicated in parentheses: GRSV-ARo (Alejandro Roca), GRSV-Baig (Cnel Baigorria), GRSV-Beng (Bengolea), GRSV-Cab (Gral Cabrera), GRSV-DVe (Dalmacio Vélez), GRSV-Etr (Etruria), GRSV-Euf (Sta Eufemia), GRSV-MGa (Monte de los Gauchos),
GRSV-Mol (Cnel Moldes), GRSV-Oli (Oliva), GRSV-RCua (Río Cuarto), GRSV-RTer (Río Tercero), GRSV-Tan (Tancacha), GRSV-TPu (Tío
Pujio), GRSV-AR (DQ973171; Argentina), GRSV-BR (AF251271; Brazil), GRSV-SA05 (S54327; South Africa), TCSV (AF282982; Brazil)
and TSWV (AB175809; Korea).
Phylogenetic analyses based on the nucleotide
sequence of the GRSV N gene showed segregation
into three clusters. The isolates from Córdoba
clustered with GRSV-AR, whereas isolates from
Brazil and South Africa clustered in separate
clades (Figure 1). This result indicates that the N gene is highly conserved and that the degree of
relatedness between GRSV isolates is correlated
with their geographic origin, which agrees with
previous studies of Tospovirus (Nischwitz et al.,
2007; Yeturu et al., 2014). This finding may be
related with the coordinated co-evolution between
thrips, tospoviruses, and their plant hosts (Ullman et al., 2005; Whitfield et al., 2005; Ananthakrishnan & Annadurai, 2007; Pappu et al., 2009; Sundaraj et
al., 2014).
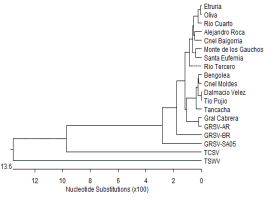
Figure 1. Phylogenetic tree showing relationships of nucleotide
sequences of the N gene between the GRSV peanut isolates from
Córdoba and reported sequences of GRSV, TCSV and TSWV in
the GenBank.
In all the evaluated fields (100%) many thrips were
detected in flowers and leaves of peanut plants.
They cause damage by inserting their needlelike
mouthparts into the leaf surface and feeding, and
according to their abundance, symptoms could be
negligible or significant. In this survey, 19 thrips
samples were taken from different locations and in
two years. A total of 1025 individuals were analyzed
(782 and 243 in the 2009 and 2010 seasons,
respectively). The most frequently observed species was F. schultzei (tomato thrips) (442 and
231 in 2009 and 1010, respectively), followed by Caliothrips phaseoli Hood (bean thrips) (321 only
in 2009) and F. occidentalis (western flower thrips),
of which only 30 thrips were collected in both years
(Table 2). Of these species, those belonging to the Frankliniella genus can transmit GRSV. F. schultzei,
the most efficient vector of GRSV (Nagata et al.,
2004), was present in almost all samples studied
both in the crop and in the associated weeds
shortpod mustard (Hirschfeldia incana (L.) Lagr.-
Fossat, family Brassicaceae), and tall morningglory (Ipomoea purpurea (L.) Roth., family Convolvulaceae), (Table 2). H. incana is an annual
to biennial plant species and has been reported as
natural host of another Tospovirus, the TSWV (Lavina
et al., 1996), so it could be a GRSV alternative host
and/or a reproductive host for thrips. F. schultzei is a
poliphagous species that is widespread throughout
Córdoba's peanut growing region and because
of the long distance movements of winged thrips
carried by winds it could be responsible for GRSV
spread. Other thrips species present in peanut
crops with high frequency was C. phaseoli. It was
detected on peanut and other widely cultivated
crops such as soybean (Glycine max (L.) Merr.)
and lucerne (Medicago sativa L.) where it causes
plant damage (Boito et al., 2006). However, it has
not been reported as a Tospovirus vector.
Table 2. Thrips species infecting peanuts and weeds sampled at different locations and on different dates in the surveyed growing area
of Córdoba province.

This work shows that peanut crops are usually
attacked by thrips. Accordingly, our study reports the presence of F. schultzei and F. occidentalis on
peanuts in Argentina for the first time. They are
important pests because besides causing leaf
damage to crops, they could be acting as vectors
of GRSV. It is likely that viruliferous F. schultzei thrips
are responsible for the dispersion of the GRSV
strain infecting the peanut crops. However, despite
the presence of the competent thrips vectors, no
plants infected with GRSV were detected in 2010,
so further investigations are needed to elucidate
the components involved in the viral disease.
Understanding the vector-virus relationship
between thrips and Tospovirus is fundamental
to develop disease management measures.
Currently, ongoing research works are focused on
the population dynamics of thrips within the peanut
fields and the role of weed host reservoirs in viral
infection cycle and virus spread.
ACKNOWLEDGEMENTS
This work was financially supported by the
Programa Nacional Cultivos Industriales-INTA and
Fundación Maní Argentino.
REFERENCES
1. Ananthakrishnan, T.N. and R.S. Annadurai, 2007. Thrips - tospovirus interactions: Biological and molecular
implications. Current Science 92: 1083-1086.
2. Boari, A.J.; E. Maciel-Zambolim, D.D. Lau, G.S.A Lima,
E.W. Kitajima, S.H Brommonschenkel and F.M.
Zerbini, 2002. Detection and partial characterization
of an isolate of Groundnut ringspot virus in Solanum
sessiliforum. Fitopatologia Brasileira 27: 249-253.
3. Boito, G.T.; J.A. Ornaghi, J.A. Giuggia and D. Giovanini,
2006. Primera cita de dos especies de insectos sobre
el cultivo de maní (Arachis hypogaea L.) en Córdoba,
Argentina. Agriscientia XXIII (2): 99-103.
4. Chu, F.-H.; C.-H. Chao, Y.-C. Peng, S.-S. Lin, C.-C. Chen
and S.-D. Yeh, 2001. Serological and molecular
characterization of Peanut chlorotic fan-spot virus, a
new species of the genus Tospovirus. Phytopathology
91: 856-863.
5. Culbreath, A.K.; J.W Todd and S.L. Brown, 2003.
Epidemiology and management of tomato spotted wilt
in peanut. Annual Review of Phytopathology 41: 53-75.
6. De Avila, A.C.; P. De Haan, R. Kormelink, R.O. Resende,
R.W. Goldbach and D. Peters, 1993. Classification of
tospoviruses based on phylogeny of nucleoprotein
gene sequences. Journal of General Virology 74: 153-159.
7. de Borbón, C.M. and O. Gracia, 1996. Frankliniella
schultzei (Tribom), eficiente vector de Groundnut
ringspot virus (GRSV) en cultivo de lechuga en
Mendoza, Argentina. Fitopalogía Brasileira 21: 423.
8. de Borbón, C.M.; O. Gracia and R. Píccolo, 2006.
Relationships between Tospovirus incidence and
thrips populations on tomato in Mendoza, Argentina.
Journal of Phytopathology 154: 93-99.
9. de Breuil, S.; J. Abad, C.F. Nome, F. Giolitti, P. López
Lambertini and S. Lenardon, 2007. Groundnut ringspot
virus: an emerging Tospovirus inducing disease in
peanut crops. Journal of Phytopathology 155: 251-
254.
10. de Breuil, S.; M.S. Nievas, F. Giolitti, L.M. Giorda and
S. Lenardon, 2008. Occurrence, prevalence and
distribution of viruses infecting peanut in Argentina.
Plant Disease 92: 1237-1240.
11. DNASTAR. 2001. Lasergene Biocomputing Software for
Windows, Ver. 5. Madison, WI.
12. King, A.M.Q.; M.J. Adams, E.B. Carstens and E.J.
Lefkowitz, 2012. Virus Taxonomy. Ninth Report of the
International Committee on Taxonomy of Virus. San
Diego, CA, Elsevier Academic Press, pp. 737-739.
13. Lavina, A.; A. Aramburu and E. Moriones, 1996.
Occurrence of tomato spotted wilt and cucumber
mosaic viruses in field grown tomato crops and
associated weeds in northeastern Spain. Plant
Pathology 45: 837-842.
14. Ministerio de Agricultura Ganadería y Pesca- Sistema
Integrado de Información Agropecuaria; Siembra,
cosecha, producción y rendimiento, 2013. <http://www.siia.gov.ar/_apps/siia/estimaciones/estima2.php> Consultada el 01/10/2014.
15. Nagata, T.; A.C.L. Almeida, R.O. Resende and A.C. de Ávila, 2004. The competence of four thrips species
to transmit and replicate four tospoviruses. Plant
Pathology 53: 136-140.
16. NCBI, 2014. National Center for Biotechnology Information. BLAST: Basic Local Alignment Search
Tool. Internet resource: http://blast.ncbi.nlm.nih.gov/.
17. Nischwitz, C.; H.R. Pappu, S.W. Mullis, A.N. Sparks,
D.B. Langston, A.S. Csinos and R.D. Gitaitis, 2007.
Phylogenetic analysis of Iris yellow spot virus isolates
from onion (Allium cepa) in Georgia (USA) and Peru.
Journal of Phytopathology 155: 531-535.
18. Pappu, H.R.; R.A.C. Jones and R.K. Jain, 2009. Global
status of tospovirus epidemics in diverse cropping
systems: Successes achieved and challenges ahead.
Virus Research 141: 219-236.
19. Reddy, D.V.R.; A.S. Ratna, M.R. Sudarshana and I. Kiran
Kumar, 1992. Serological relationships and purification
of bud necrosis virus, a tospovirus occurring in peanut
(Arachis hypogaea L.) in India. Annals of Applied
Biology 120: 279-286.
20. Satyanarayana, T.; S. Gowda, K. Lakshminarayana
Reddy, S.E. Mitcell, W.O. Dawson and D.V.R. Reddy,
1998. Peanut yellow spot virus is a member of a new
serogroup of Tospovirus genus based on small (S)
RNA sequence and organization. Archives of Virology
143: 353-364.
21. Sundaraj, S.; R. Srinivasan, A.K. Culbreath, D.G. Riley
and H.R. Pappu, 2014. Host plant resistance against Tomato spotted wilt virus in peanut (Arachis hypogaea)
and its impact on susceptibility to the virus, virus
population genetics, and vector feeding behavior and
survival. Phytopathology 104: 202-210.
22. Ullman, D.E.; A.E. Whitfield and T.L. German, 2005. Thrips
and tospoviruses come of age: Mapping determinants
of insect transmission. Proceedings of the National
Academy of Sciences, USA 102: 4931-32.
23. Whitfeld, A.E.; D.E. Ullman and T.L. German, 2005.
Tospovirus-thrips interactions. Annual Review of
Phytopathology 43: 459-489.
24. Yeturu, S.; B.V. Bhaskara Reddy, S. Asadhi and D.V.R Sai
Gopal, 2014. Sequence diversity of the nucleoprotein
gene of Peanut bud necrosis isolates from South India.
Journal of Phytopathology 162: 542-547.


