Carbapenem-resistant Klebsiella pneumoniae (CR-Kp) infections are associated with high morbidity and mortality rates, mainly in patients admitted to intensive care units. Nowadays, the production of KPC-p-lactamases con-stitutes the main resistance mechanism among CR-Kp. KPC-producing K. pneumoniae (KPC-Kp) became endemic in 2010 in Argentinian hospitals, and has been attributed to the global expansion of the epidemic clone K. pneumoniae sequence type (ST) 258 and related strains of the clonal complex (CC) 2584. However, in recent years, the emergence of KPC-Kp belonging to non-CC258 has been reported3, including the high-risk clone K. pneumoniae ST307. Although ST307 was reported in several countries as early as 2013, it emerged in 1994, close to the estimated emergence of the international ST258 clone15. ST307 was initially associated with blaCTX-M-15; however, enough indexed literature supports that this lineage could acquire and disseminate carbapene-mases (blaKPC, blaNDM, blaoxA-48, blaoxA-48, blaoxA-181, blaGEs-5) and probably other clinically important antimicrobial resistance determinants14,15. K. pneumoniae ST307 is a highly successful MDR clone that has been proposed as a candidate for becoming the prevalent high-risk CR-Kp clone based on its greater aptitude, persistence and adaptation to hospital environments and to the human host14. K. pneumoniae ST307 was first noticed in Argentina during a surveillance study performed in 2015-2017. Two out of 76 KPC-Kp stud-ied isolates, named Kp2 and Kp14, belonged to ST307. Kp2 corresponded to a KPC-2-producing K. pneumoniae isolate, which was recovered in January 2017, in a hospital located in Buenos Aires (HA), being the first KPC-Kp ST307 reported in Argentina. Kp14 was the first KPC-3-producing K. pneu-moniae ST307 identified in this country, and corresponded to the first detection of this KPC allele3. Kp14 was isolated in December 2017 in another hospital in Buenos Aires (HB). The aim of this study was to investigate the presence of CR-Kp ST307 isolates and to characterize them based on their resistance mechanisms and clonal relationship.
From November 2018 to February 2019, a total of 35 CR-Kp isolates were recovered from inpatients with invasive infections in HB. These isolates were anonymously deliv-ered to the Instituto de Investigaciones en Bacteriología y Virología Molecular (IBaViM) - FFyB, Universidad de Buenos Aires for further characterization. Those previously men-tioned K. pneumoniae ST307 isolates, Kp2 and Kp14, were included in this study. Except for Kp2, the isolates were recovered in HB. Susceptibility to p-lactams, quinolones, aminoglycosides and trimethoprim-sulfamethoxazole (TMS) was assessed by automated systems (BD Phoenix). Aztreonam and ceftazidime/abivactam (CZA) susceptibilities were determined by the disk diffusion method. Antimi-crobial susceptibilities were interpreted according to CLSI M100-ED29 2019 (https://clsi.org/all-free-resources/). Col-istin minimum inhibitory concentration was determined by the broth microdilution method according to EUCAST 2019 (http://www.eucast.org/). Double disk synergy tests were performed in order to detect KPC and/or metallo-p-lactamases (MBL) using phenyl boronic acid (300 ^g) and EDTA (1 ^mol) disks, respectively9,11.
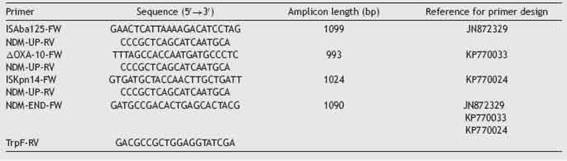
The presence of the most prevalent carbapenemase encoding genes (blavIM, bla^P, blasPM, blaKPC, blaNDM and blaOXA-48-like) was investigated by multiplex-PCR, using specific primers and total DNA as template13. More-over, blaKPC and blaNDM were confirmed as previously described using plasmid extracted DNA as template1,12. PCR products were sequenced at external facilities (Macrogen, Korea) and analyzed using the BLAST tool at the National Center for Biotechnology Information (http://www.ncbi.nlm.nih.gov/BLAST). The genetic con-text of blaKPC was studied by PCR mapping and sequencing in order to infer the Tn4401 structures or blaKPC-bearing non-Tn4401 elements.3 To investigate blaNDM-1 surrounding regions, specific primers were designed based on previously reported structures (Table 1). In those isolates that displayed colistin resistance, screening for plasmid located colistin resistance genes (mcr-1,-2,-3,-4,-5) was conducted by multiplex PCR as previously described.7 Plasmid conju-gation assays were performed in Luria Bertani broth (LB) using E. coli J53AzR as recipient strain and K. pneumoniae isolates as the donor strains. Isolates were grown overnight in LB (5ml) at 37°C. Afterincubation, the bacterial cultures were mixed 1:10 and 5:10 donor-recipient, and incubated at 37°C for 18 h without shaking. Transconjugants were selected on Mueller Hinton agar plates containing sodium azide (200 ^g/ml) and imipenem (0.5 ^g/ml). Plasmidsfrom transconjugants were extracted and further characterized in terms of their replicons2,13. Plasmid size was estimated by treatment with S1 nuclease followed by pulse-field gel electrophoresis (PFGE).

Table 2: Characteristics of carbapenem-resistant Klebsiella pneumoniae ST307 isolates included in this study.
Multilocus sequence typing was performed amplifying and sequencing the following seven house-keeping genes: gapA, infB, mdh, pgi, phoE, rpoB, tonB, in accordance with www.pubmlst.org. Clonal relationship was analyzed by PFGE after digestion of genomic DNA with XbaI. Among the 35 CR-Kp isolates, 4 were assigned to ST307 according to www.pubmlst.org. These CR-Kp ST307 isolates, KpM24, KpM25, KpM30, KpM34 and those previously recovered CR-Kp, Kp2 and Kp14, were further characterized. All of them displayed resistance to ampicillin, ampicillin-sulbactam, piperacillin-tazobactam, ceftazidime, ceftriaxone, aztre-onam, imipenem, meropenem, ciprofloxacin, levofloxacin and gentamicin. KpM24, KpM25 and KpM34 isolates were resistant to CZA, while Kp2, Kp14 and KpM30 remained susceptible. Except for Kp2, they also displayed resistance to colistin (Table 2). Phenotypic screening of KPC enzymes rendered positive results in Kp2, Kp14 and KpM30, while the production of carbapenemases could not be phenotypically detected in the remaining isolates.
The presence of blaKPC was detected in all CR-Kp ST307 isolates, corresponding to either blaKPC-2 or blaKPC-3. KpM24, KpM25 and KpM34 harbored both blaKPC-3 and blaNDM-i (Table 2). Tn4401a was recognized as the genetic platform for blaKPC-2 and blaKPC-3. Moreover, the presence of ISKpn14 was detected in the upstream region of blaNDM-1, followed by a partial sequence of ISAba125, while bleMBL and trpF were detected downstream. blaNDM-1 harboringplasmidsweresuc-cessfully transferred by conjugation to an Escherichia coli J53 recipient strain, and all blaNDM-1 plasmids from transcon-jugants corresponded to the A/C incompatibility group and were approximately 145.5 kb in size. blaKPC harboring plas-mids could not be transferred under the assayed conditions. Screening for plasmid located colistin resistance genes ren-dered negative results.
KpM24, KpM25, KpM30 and Kp14 displayed a unique pulse-type in the XbaI-PFGE, closely related to KpM34 (Fig. 1). As previously mentioned, all these isolates were recovered in HB. Kp2, which was recovered in HA, exhibited a dif-ferent pulsetype. An increase of K. pneumoniae ST307 was observed in HB compared with previous periods3, despite the fact that a longer time should be monitored for a conclu-sive epidemiological definition. ST307 has become a globally
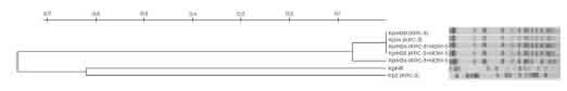
Figure 1: Clonal relationship of carbapenemase-producing K. pneumoniae ST307 isolates. Footnote: KpNR corresponds to a non-related KPC-2 producing isolate.
In conclusion, the present study provides information about the dissemination of this high-risk clone in HB, although it is still infrequently reported in Argentina. Here we alert about the emergence of K. pneumoniae ST307 iso-lates co-producing KPC-3 and NDM-1, which could lead to a worrisome scenario in this country, regarding the remark-able features of K. pneumoniae ST307 and its resistance profile to CZA. Association of blaNDM-1 to epidemic plasmids, such as the IncA/C group, along with the introduction of avibactam in HB in 2018, may have favored the selection and dissemination of NDM-1-producing isolates. The potential for dissemination of carbapenemase-producing K. pneumoniae ST307 warrants larger and more continuous epidemiologi-cal surveillance studies aimed at the early detection of this high-risk clone.
FundingThis work was supported by UBACyT to M. Radice and G. Gutkind (20020150100174BA and 20020170100473BA, respectively); PICT to G. Gutkind (2015-1925); PIP to M. Radice (PIP11220200102425CO).
Ethical approvalThe ethics committee of FFyB-UBA approved this study (Res CD 894-2019). All isolates were delivered anonymized from HB to IBaViM-FFyB-UBA, in order to preserve patient’s identity. Isolates have been designated using numbers, and hospitals were mentioned using capital letters.












 uBio
uBio 


