INTRODUCTION
Acquired Immunodeficiency Syndrome (AIDS) is characterized by severe immunologic alterations mainly affecting CD4+ T cells, a subpopulation of T lymphocytes, and by subsequent development of potentially fatal opportunistic infections.
According to the World Health Organization, approximately 34 million people are infected with human immunodeficiency virus (HIV), and although there has been much progress in HIV research in recent years, including therapies such as highly active antiretroviral therapy (HAART), this disease still constitutes one of the most significant public health problems in the world1. The National Ministry of Health estimates that in Argentina there are 120,000 people living with human immunodeficiency virus, 30% of whom are unaware of their HIV status2.
Since the introduction of HAART (highly active antiretroviral therapy), the epidemiological profiles of the link between infectious disease and HIV have changed3,4. The factors involved in these changes include partial restoration of the immune system, control of viral load and stability of the immune system over time5. HIV infection has turned into a chronic disease. Global mortality rates decreased from 97.4% in 1993 to 19.8% in 20016. The current guidelines for antiretroviral therapy recommend initiating treatment earlier7.
However, the disease has not yet been completely eradicated, probably due to the ability of the virus to remain latent in host immune cells (T lymphocytes, macrophages and dendritic cells), as well as to its high mutation rate8.
Periodontitis, as a multifactorial ecological disease whose primary aetiology is bacterial biofilm, causes a chronic inflammatory response by the release of bacterial and host cell products9. Bacteria from subgingival periodontopathic biofilm have previously been shown to be a significant risk factor for many systemic diseases (heart disease, diabetes and low birth-weight, among others)10.
Epidemiological studies that used continuous measures of probing depth and clinical attachment loss have shown that advanced forms of periodontitis that resulting in severe loss of supporting structures and substantial tooth loss affect 10-15% of the population globally11. This estimated prevalence range includes severe periodontitis, which primarily affects adolescents, young adults and adults, and whose prevalence increases with age in all populations12-14.
Longitudinal studies have shown that prolonged use of HAART in HIV-infected periodontal patients was associated with a decrease in the prevalence of putative periodontal pathogens as compared to seronegative patients. However, seropositive patients had species not usually associated with periodontitis, regardless of their periodontal status15. Establishing comparisons among studies conducted in the pre-and post HAART eras is complex given the need for appropriate and consistent characterization of HIV-associated periodontal disease to avoid underestimating the prevalence and severity of the infection16. The impact of HAART on periodontal disease in HIV patients in developing countries such as Argentina remains unclear, and reports on periodontal microbiota in this population are not conclusive.
Recent studies suggest that bacteria found in the oral cavity, including P. gingivalis, are able to reactivate the latent HIV virus within infected cells. In 2009, Huang et al. evaluated the capacity of bacteria found in the oral cavity to stimulate HIV promoter activation17.
Research has been conducted in which cellular T-lymphocyte, macrophage and fibroblast lines transfected with the HIV long terminal repeat promoter were stimulated with oral bacterial sonicates, and both the Gram-negative and Gram-positive species were able to elicit positive stimulatory activity18. Another assay suggests that P gingivalis produces a fatty acid called butyric acid, which may induce reactivation of the latent HIV-1 virus9. González et al. looked specifically at the ability of supernatants (cytokines/chemokines) produced by human gingival fibroblasts and oral epithelial cells to modulate HIV promoter activation in macrophages when challenged with periodontal pathogens. Stimulation with P. gingivalis and extract of F. nucleatum showed a significant increase in cytokines/chemokines, especially Interleukin-6 and Interleukin-8, which have the ability to modulate the activation of HIV-1 promoters18.
The superposition of events such as these, with reactivation of the HIV virus due to the coexistence of active periodontal disease, increase the systemic inflammatory state of the host. This worsens the effector response and subsequent clinical outcome19. The aim of this study was to describe the microbiological profile of periodontal pathogens in a population of local patients living with HIV/AIDS on highly active antiretroviral therapy.
MATERIALS AND METHODS
The population study comprised male and female HIV-infected patients treated with HAART who received care the High-Risk Patients Dental Care Unit (CLAPAR I), School of Dentistry, University of Buenos Aires, Argentina. The patients were informed of the study objectives, risks and benefits. All participants agreed to participate voluntarily and signed an informed consent form approved by the Ethics Committee of the School of Dentistry, University of Buenos Aires. This study fulfils the requirements of STROBE rules for human observational studies.
This descriptive study focused on people between 21 and 65 years old living with HIV/AIDS who had been under treatment with HAART for at least six months prior to the dental examination. A nonprobabilistic sequential sample of 50 subjects seeking care at the CLAPAR I was evaluated. Information on the patients' medical history was obtained from clinical records (antiretroviral treatment history, viral load and CD4+ levels). Viral load was detected by real-time quantitative polymerase chain reaction (qPCR) to quantify plasma HIV-1 RNA, using ABBOTT REAL TIME HIV-1 RNA VERSION 3.0 with a sensitivity of 40 HIV-1RNA copies/ml (a viral load below 50 copies/ml was considered undetectable). The missing data from the clinical histories were traced in each of the hospital's services.
Dental clinical examination was performed by a calibrated researcher (intra-examiner kappa >0.80) using standard dental instruments (Marquis periodontal probe) to determine probing depth (PD), clinical attachment level (CAL) and bleeding on probing (BOP). Clinical parameters were evaluated at 4,765 periodontal sites, 8 sites per tooth, in order to provide more accuracy for the description of periodontal status.
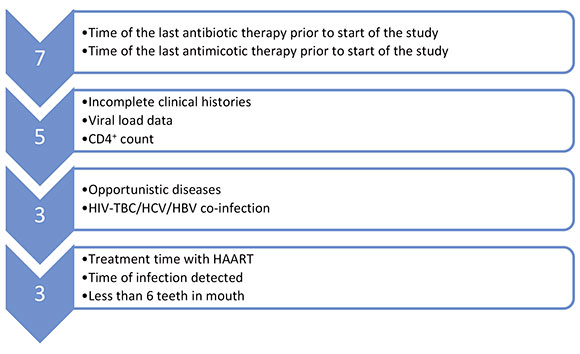
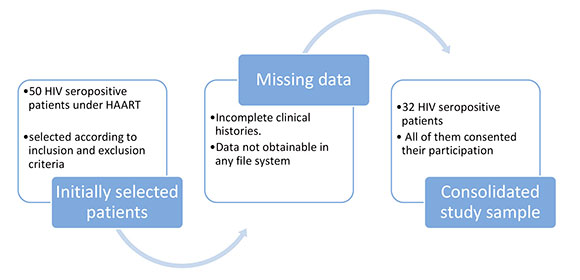
A total 50 persons were given appointments for screening for eligibility. Exclusion criteria were: less than 6 teeth (n=3), antibiotic or antifungal treatment within the past 6 months (n=7), incomplete clinical histories, CD4+ lymphocyte level count lower than 250 cells/ml and absence of viral load data (n = 5), concomitant systemic or infectious diseases (n = 3). As a result of the selection, 32 patients were included in the study protocol. All evaluated patients received their dental treatment and follow-up in CLAPAR 1 (Figs. 1 and 2).
Based on the most severe gingival inflammatory signs, one site per quadrant was selected to be studied microbiologically (n=128 sites). Samples of subgingival biofilm were collected by absorption using 4 sterile N°35 paper points. They were placed one at a time into the depth of the pocket and removed after 20 seconds. The 4 paper points from each site were placed in 1ml of refrigerated transport fluid (RTF)20. Smears were prepared with the material obtained from the soft wall of each periodontal site.
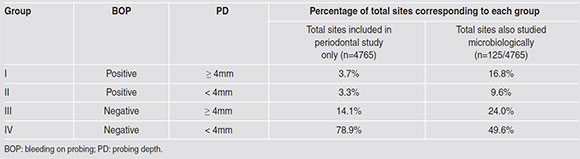
For the microbiological assay, the periodontal pockets studied were assigned to four groups (Groups I to IV) according to their periodontal parameters such as bleeding on probing (BOP) and probing depth (PD). Group I: BOP/PD > 4mm n=21 (16.4 %); Group II: BOP/PD < 4mm n=30 (23.4%); Group III: NO BOP/PD > 4mm n=12 (9.4%); Group IV: NO BOP/PD <4mm n=62 (49.6%). Details are provided in Table 3.
Microbiological Processing
To isolate strict anaerobic microorganisms, 100 pl of the homogenized diluted (10-2, 10-3) samples were cultured in Anaerobic Blood Agar (ABA-BHI Agar) supplemented with 5% lacked sheep blood (Laboratorio Gutiérrez, Argentina), 5 pg/ml of hemin (Sigma-Aldrich®, Argentina),and 1 pg/ml of menadione (Sigma-Aldrich®, Saint Louis, MO, USA). The plates were incubated under strict anaerobic conditions, and analysed at 7 and 15 days (GasPakTM, Mitsubishi, Medica-tec®). For detection of Aggregatibacter actinomycetem comitans (Aa), enteric bacteria, non-fermenting gram negative bacilli (NFGNB), Staphylococcus aureus (Sa), coagulase-negative staphylococci (CoNS), and Candida spp, 100 pl of the samples were cultured directly on selective and differential agar medium as follows: AASM21 for Aa, Levine (Oxoid®, UK) and CLDE (Oxoid®, UK) for enteric bacteria, bile esculin agar (Britania Laboratory, Argentina) supplemented with 0.04% potassium tellurite for enterococci, Chapman's mannitol salt agar (Oxoid®, UK) for Sa, and CHROMagar® Candida for Candida spp. The plates were incubated under aerobic or capnophilic conditions according to bacterial atmospheric requirements, for 24h, 48h and 5 days. In all cases, the species were biotyped using conventional methods (Rosco® Diatabs) and a commercially available kit (Remel® enteric bacteria).
Molecular processing
For molecular processing, 200ul of each sample were stored at -20°C, and processed using a commercially available kit (Bioneer® Genbiotech) for extraction of genomic DNA. Seven "in house" end point polymerase chain reactions (PCR) were run for each sample (n=896), using specific primers (Ashimoto et al. 1996) for detecting periodon -topathic bacteria of the red complex (RCB) i.e. Porphyromonas gingivalis (Pg), Treponema denticola (Td), and Tannerella forsythia (Tf), and of the orange complex (OCB), i.e. Fusobacterium nucleatum (Fn), Prevotella intermedia (Pi) and Aggregatibacter actinomycetem comitans (Aa).
The amplification products were separated by electrophoresis in agarose gels (2%) in TAE buffer22, stained with Gold View (Montebio), and analysed using the Gel DocTM XR+ Imaging System (Bio-rad). Negative and positive controls were used in all cases (Porphyromonas gingivalis ATCC 33277, Prevotella intermedia ATCC 25611, Fusobacterium nucleatum ATCC 10953, Aggregatibacter actinomycetamcomitans ATCC 29522, Tannerella forsythia ATCC 43037), and DNA of wild-type strains, which were characterized, biotyped, and stored in the Collection of Microbial Cultures (Laboratory for Microbiological Diagnosis of the School of Dentistry, University of Buenos Aires, Argentina). Ubiquitous (Ubi) primers were also used to detect genomic bacterial DNA.
Statistical Analysis
Quantitative variables are expressed as average, standard deviation, median, and percentiles. Comparison of quantitative variables and factors or groups was performed using Student's t-test for independent samples. In all cases, homoscedasticity was evaluated using Levene's Test, and data normality was analysed using Kolmogorov Smirnov and Shapiro Wilks tests, as well as graphic normality tests. The data that did not meet parametric test assumptions were analysed using Kruskal Wallis test and post hoc Dunn-Bonferroni contrast test. Categorical variables were analysed and compared using Chi-square test and Yates continuity corrections.
When distribution of data was binomial, percentages were compared using the proportions test (difference of percentages) and a normal approximation. Comparisons among more than two groups were performed applying a Bonferroni correction to the level of significance.
In all cases, the level of significance was set at 5% to rule out the null hypothesis. All statistical analyses were performed using statistical software SPSS (version 25), STATA 14, MS Excel 2016, Epidat 4.2.
RESULTS
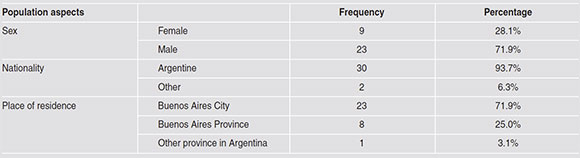
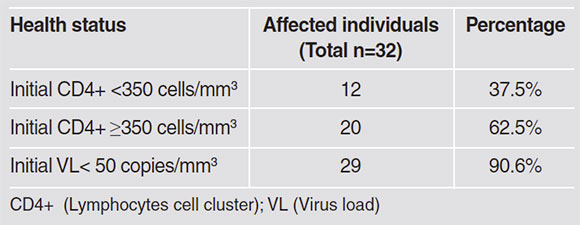
The study comprised 32 patients (9 female and 23 male). All participants fulfilled the inclusion criteria, as shown in Flowcharts 1 and 2. Of the study patients, who were mostly Argentinians, 90.6% had less than 50 copies/mm3 viral load; data shown in Tables 1 and 2. The proportion of active periodontal sites and their distribution according to the analysis groups are shown in Table 3.
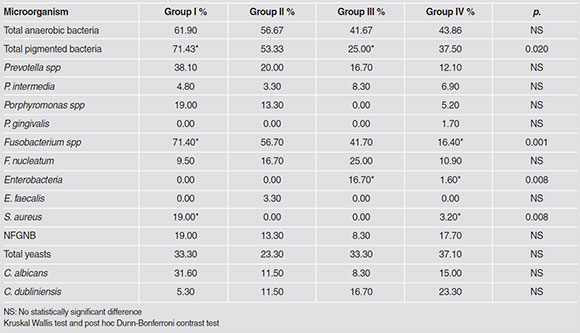
Isolates of conventional microorganisms were headed by Candida spp. (32.8%; C. albicans: 15.2%; C. dubliniensis: 16%) closely followed by Gram-negative bacilli (18.4%; NFGNB: 16%; enteric bacteria: 2.4%), Staphylococcus spp. 17.6% (CoNS 12.8%; S. aureus: 4.8%), and Enterococcus faecalis (0.8%) in the last position. Recovery rates of microorganisms using conventional cultures, and comparison of recovery rates among groups are shown in Table 4.
The mean anaerobic bacterial count (CFU) was 1.45 x 106 CFU/ml, and there were significant differences in total counts of pigmented anaerobic and fusiform bacteria between Groups I and IV, as expected.
Conventional anaerobic culture and biotyping methods were unable to establish significant differences in the prevalence of P.intermedia and P. gingivalis in any of the study groups. Similarly, different Fusobacterium species were observed but could not be identified at the species/subspecies level in any of the cases except as F. nucleatum. Microbiological results for opportunistic bacteria such as S. aureus showed statistically significant differences in Group I versus the others (p<0. 01). Enteric bacteria were only isolated from Groups III and IV. No statistically significant difference was found for recovery rates for E. faecalis, NFGNB or total yeasts between the study groups.
Endpoint PCR results established statistically significant differences between groups according to periodontal clinical parameters (GI/GIV). P.gingivalis, T.denticola and Tforsythia, as periodontopathic species, were the most prevalent (66.7% Pg, Tf 52.4% Td) in the group of patients with active periodontal parameters (GI) (Table 5).
DISCUSSION
Studies reported in the literature have shown a similar composition of subgingival microbiota in seronegative and seropositive HIV patients23,24,15. More recently, in 2019, a study on the subgingival periodontopathic microbiome showed that although P.gingivalis is the marker species of the "red complex", Treponema denticola and Tanerella forsythia as well as Filifactor alocis, Mitsuokella dentalis, Slackia exigua, Dialister pneumosintes, Selenomonas sputigena and Treponema lecithinolyticum are among the ten best biomarkers for this pathology25. However, other reports26,27 have found seropositive HIV patients to show a higher prevalence of putative periodontal pathogens such as A. actinomycetemcomitans spp., P. intermedia and T. denticola, as well as combinations of species. In addition, microbiological studies have reported isolation of species not usually associated with chronic periodontitis, including Staphylococcus epidermidis, C. albicans, E. faecalis, Clostridium difficile, Clostridium clostridiforme, Klebsiella pneumoniae and Mycoplasma salivarius, in subgingival biofilm samples from HIV-infected patients28-31. Aas et al. studied the subgingival biofilm of seropositive patients with different immune status32. The authors found microorganisms such as Saccharomyces cerevisiae in subjects with higher viral load and low lymphocyte counts, lending support to the opportunistic nature of periodontal infections in these patients. Gonçalves et al. found higher prevalence of Enterococcus faecalis and Acinetobacter baumanii in subgingival biofilm of HIV+ patients on HAART with CD4+ T-lymphocyte counts < 200 cells/mm3 and viral loads in the range of 105 copies/ml compared to seronegative patients, regardless of their periodontal status15. Conversely, recovery of opportunistic microorganisms was not significant in our study population, in which 90.6% of the participants showed undetectable viral loads, and 62.5% had CD4+ T lymphocyte counts above 350 cells/mm3. These discrepancies may be due to differences in the immune status of the study populations. Gonsalves et al. observed a lower incidence of periodontal lesions in patients under HAART33. Despite the lower frequency of clinical determinants of periodontal disease in our study population, more than 50% of the study sites in Group 1 were positive for periodontopathic bacteria belonging to Socransky's red complex, which were detected using endpoint PCR34. The primers designed by Ashimoto enable detection of A. actinomycetemcomitans, P. gingivalis, P. intermedia, T. denticola and T. forsythia DNA in subgingival biofilm samples with high sensitivity and specificity (more than 99.9%). These primers have been used successfully by different authors35 and therefore we consider them reliable for use in our study.
Brenchley et al. and Collman et al. posited that infections by Gram-negative bacteria and bacterial translocation associated with the mucosa in HIV-1-infected patients might directly or indirectly contribute to virus reactivation via pro-inflammatory cytokines released from stimulated cells36,37. Periodontal disease, seen as a chronic polymicrobial, ecological and immune-inflammatory disease, could play a role in viral reactivation in HIV seropositive patients38.
Because HIV infection affects the host's immune response39, gaining better understanding of the role of gingival-periodontal disease in immune activation in HIV seropositive patients would seem to be of the utmost importance for the development of therapeutic strategies aimed at controlling virus reactivation, patient immune status, and progression of AIDS.
Patients living with HIV/AIDS on HAART that were studied together in the High-Risk Patients Dental Care Unit (CLAPAR I) and the Micro -biological Diagnostic Laboratory at School of Dentistry, University of Buenos Aires, Argentina showed low prevalence (1%) of periodontal lesions. Yeast and non-fermenting Gram-negative bacilli were distributed homogeneously, regardless of the depth of the periodontal pocket and in quantities that resemble a colonizing function.
Despite the low prevalence of periodontal disease and the moderate level of activity recorded in this study, P. gingivalis, T. denticola and T. fosythia are within the framework of the microbiological profile of the subgingival biofilm of the eligible HIV-seropositive population. They were detected more frequently in periodontal active pockets (GI). As known, these pathogens and their products can contribute to systemic inflammation, so periodontal treatment could contribute to overcoming, and there by improving the general condition of seropositive HIV patients.
This is the first local study to record the prevalence of periodontopathic microorganisms in periodontal pockets of HIV patients. The results justify further in-depth longitudinal studies of the subgingival microbiome of local patients living with HIV/AIDS who are on HAART, which might clarify many of the interactions of these and other still unknown periodontal bacteria, describe new pathogenicity factors and contribute to understanding the particular dynamics of these subgingival microbiomes and HIV-1 virus homeostasis.