INTRODUCTION
The genus Coniophora (Coniophoraceae, Boletales) is a widespread brown-rot wood decay fungus occurring in building timber as well as in natural environments, mostly in temperate regions (Kauserud et al., 2007). Coniophora arida and C. olivacea have been recorded on dead wood of diverse tree species, both conifer and hardwood tim- ber, and are distributed in northern and southern temperate zones worldwide. Conio- phora olivacea has also been reported on living Picea spp. in North America. Both C. arida and C. olivacea are common in coniferous forests (Ginns, 1982). Together with
C. puteana, they cause economically important losses in several ways; they frequently cause decay of living trees and have been observed indoors on construction materials (Kauserud et al., 2007). Chee et al. (1998) report significant weight losses of wood in solid-state cultures of C. olivacea. Da Costa et al. (1967) also mention the capability of vigorous attack of this species on a very wide range of timbers under different uses, including hardwood trees such as Eucalyptus marginata in Western Australia. They also report that wood density was significantly affected by Coniophora olivacea whereas other fungal species tested did not significantly alter this parameter. Rud- man (1963) proves that the sapwood of many tree species is uniformly susceptible to attack by this fungus. Although the infective role of each dissemination form is yet unknown (Edman & Jonsson, 2001), C. olivacea is able to produce basidiospores as well as mycelial cords (Stalpers, 1978), which suggests the possibility of infection through the soil from one log to another.
A specimen collected by the authors identified as C. olivacea and deposited at the CORD Herbarium (CORD FK16004) is to our knowledge the first and only record of this species in southern South America. Since the Andean Patagonia has a long history of cultivation of non-native trees species (e.g. Pinus spp., Eucalyptus spp., Pseudotsuga menziesii, among others), events of introduction of exotic fungal species are likely. Moreover, intense fieldwork performed in the last decades by many mycologists aiming to assess in depth the aphyllophoroid funga of the region did not detect this species (e.g., Greslebin & Rajchenberg, 2003).
Our collection consists of a small decayed piece of wood found on soil in a mixed forest. Given the fragmentary nature of the material, the identity of the wood species of the substrate is not obvious, and intensive histochemical work is needed to accurately identify it.
Since C. olivacea has been shown to be capable of producing significant damage in wooden structures, reliable information is needed as a base for future management programs. Strategies aiming to restrict the dispersion of forest pathogens, such as Phytophthora austrocedri, have been already promoted (Giordana et al., 2020), includ- ing restrictions in tourism and animal farming in certain areas or even chemical treatments (Silva et al., 2016). Initiatives against the spread of C. olivacea as well as against other non-native species might be needed in the future, even if they do not mainly affect living trees, since wood is the dominant construction material in the region, especially in the Chilean slope of the Andes.
Given the complex and multicausal nature of biological invasions, it is difficult to predict the behavior of a newly arrived alien species in a given area is, at least, challenging (Desprez-Loustau et al., 2007), especially when this behavior is also linked to plant species that might act as hosts, partners or reservoirs of the fungal entity (Dickie et al., 2017). However, methods taking on account climatic and other variables produce robust models outlining the potential distribution of a species given their environmental preferences. In this line, Maximum Entropy (Maxent) models have been shown to be efficient in estimating the potential geographic dis- tributions of species by merging presence-only data with environmental information (Phillips et al., 2006; Yuan et al., 2015).
An additional challenge regarding the identification of C. olivacea is the pres- ence of distinct evolutionary clades within this species (Kauserud et al., 2007). Since this diversity is morphologically cryptic, only molecular methodologies can place a specimen in this scheme. Studies taking into account this hidden diversity and aiming to find different behaviors (e.g. in degradation, dispersal, infectivity) need available material associated with reliable information on the specimen.
The aim of this work is to provide reliable information on the identity, host and potential distribution of C. olivacea, reported here as new for Patagonia, so future decisions on non-native species management can be made on a solid basis.
MATERIALS AND METHODS
Sequences and molecular analyses
DNA of the vouchered specimen was extracted with the GenElute Plant Genomic DNA Miniprep Kit (SigmaAldrich) following the manufacturer’s instructions. The rDNA ITS1-5.8S-ITS2 region (ITS) was amplified using the primers ITS1F/ITS4 White et al., 1990; Gardes & Bruns, 1993). The PCR reaction was performed with the REDExtract-N-Amp PCR ReadyMix (SigmaAldrich) for both regions under the following conditions: an initial denaturing step at 95 ºC for 4 min, 36 cycles of 30 s at 94 ºC, 45 s at 55 ºC and 70 s at 72 ºC and a final extension for 10 min at 72 ºC. The amplified DNA fragment was purified and sequenced in Macrogen (Seoul, South Korea) using the primer ITS4 for ITS. This newly generated sequence was upload- ed to GenBank with the accession number ON832657. Internal Transcribed Spacer sequences of the most similar taxa identified to species were queried in GenBank NCBI (http://www.ncbi.nlm.gov) using the MEGABLAST option (see Table S1). Sequences of specimens of the genus Leucogyrophana were recovered from GenBank and used to root the tree.
The sequences were aligned with L-INS-i strategy as implemented in MAFFT
7.0 (Katoh & Standley, 2013) and analyzed without additional editing. Nucleotide substitution models were chosen with jModelTest 2.1 (Darriba et al., 2012), under the Akaike information criterion (AIC). Maximum likelihood (ML) analysis was performed in PHYML (Guindon et al., 2010). Bootstrap values of the most likely tree were calculated after 1,000 repetitions with 10 random addition sequences and the topologies were produced using TBR swapping. B/MCMC analyses were conducted with MrBayes (Huelsenbeck & Ronquist, 2001) with 7,000,000 generations starting with a random tree and running four simultaneous chains. The first 100,000 gen- erations were discarded as burn-in. TRACER1 (http://evolve.zoo.ox.ac.uk/software. html/tracer/) was used to verify that stationarity was achieved after the first 100,000 generations.
Wood identification and characterization of the damage
Transverse, tangential and radial sections (10 μm thick) were made using a Leica Hn 40 sledge microtome and stained with cotton blue, after bleaching the wood with hydrogen peroxide 30% for 1.30 h, rinsing with distilled water 3 to 4 times, and rinsing before mounting in lactoglycerol. Ten slides of each sample were mounted and 10 unstained slides were prepared as control. Observations were made under a light microscope Leica DM500 and photomicrographs were obtained with a Canon EOS Rebel T3i digital camera. Wood library samples and the InsideWood database (http://insidewood.lib.ncsu.edu) were used for comparative study and to determine the species identity of the wood substrate.
The slides (at least 5 for each section of the procedures of noninfected and infected wood) were stained using specific procedures mentioned by Krishnamur- thy (1988) for localizing lignin, carbohydrates, and other phenolics. Lignin was revealed as red using Wiesner’s reagent for five minutes and mounted on a drop of 6N HCl. Toluidine blue was used to contrast lignin and polyphenols (turquoise green), carbohydrates (lilac) and oils (without color), and rinsed with water (Rivera Nava et al., 1999).
Occurrence data, climatic data, and niche modeling
Ecological niche model (ENM) was employed to infer the potential distribution of C. olivacea in the world, with a special focus on Patagonia. Geographical coor- dinates of curated registers of this species were recorded from Global Biodiversity Information Facility (GBIF website: www.gbif.org) and also from BLAST queries of ITS sequences in public databases included in the phylogenetic analyses (Table S2). Records before 1950 were eliminated since climatic data before is not system- atically curated. Records without documented authoritative identification were also discarded. To avoid an autocorrelation spatial effect in the ENM, the points located closer than 50 km from each other were eliminated using the R package “Wallace” v1.0.6.1 (Kass et al., 2018) implemented in R v3.6.1. The final database contained 113 records distributed on every continent except Antarctica (Table S2).
A set of 19 bioclimatic variables layers and one elevation layer were obtained from the WorldClim 2 database (http://www.worldclim.org) at 2.5 arc-min (i.e. ca. 5 km) resolution covering the entire planet (Hijmans et al., 2005). In order to minimize biased fitting of the models produced by the covariance between variables, Pearson’s correlation was performed to eliminate correlated environmental and elevation vari- ables using R v3.6.1. After we excluded correlated variables (r > 0.7), we selected six climatic variables (Fig. S1): Bio1 (annual mean temperature), Bio2 (mean diurnal range), Bio4 (annual temperature range), Bio11 (precipitation of driest month), Bio12 (annual precipitation), Bio15 (precipitation seasonality) and the elevation variable. MaxEnt was run under default settings, including auto-features, a maximum of 500 iterations, 100 replicates to ensure reliable results, and 10,000 randomly sampled background points, with a regularization multiplier of 1 and a convergence threshold of 10-5. K-fold cross-validation was used to test the performance of the model. To do this, occurrence data were randomly split into a number of equal-size groups, and models are created after resampling the set. The left-out folds were then used for evaluation. Logistic output models were produced and uploaded to ASCII in raster format and then imported to the QGIS “Geographic Information System” Version
3.4.12 (2019). The average model was geographically projected dividing the proba-
bility of occurrence into five categories: values below the threshold were considered absent, “low” threshold-0.25, “medium” 0.25-0.5, “high” 0.5-0.75, and “very high”
> 0.75. The area under the “receiver operating characteristic (ROC) curve” (AUC) values (Peterson et al., 2008; Elith & Leathwick, 2009) were used to evaluate the fitness of the prediction. Variable importance to ENMs was evaluated based on the permutation value, percent contribution, jackknife resampling, and the response curves produced by MaxEnt (Phillips et al., 2017).
RESULTS
Molecular identification
The analyses performed to support the morphological identification of the voucher FK16004 as Coniophora olivacea (Fr.) P. Karst. (Fig. 1) and also strongly suggest that this material is affiliated with group 4 within the scheme proposed by Kauserud et al. (2007). This group consists of materials found in British Columbia (Canada) and the UK, and Kauserud et al. (2007) conclude that this lineage probably has a wider boreal distribution. But more profound studies and a wider sampling is needed before proposing these localities as the origin of the introduced strain. Group 5 has not been recovered from the dataset since no available sequences representing it were retrieved from public databases.
Identification of the substrate and effect of the degradation
The woody substrate presented the following characteristics: In cross-section, dif- fuse porosity was observed, with solitary and grouped vessels colonized by abun- dant hyphae. Sectors without fibers or parenchymal rays (disintegrated tissue) were observed. In the radial section, rays of reddish coloration and with contents were
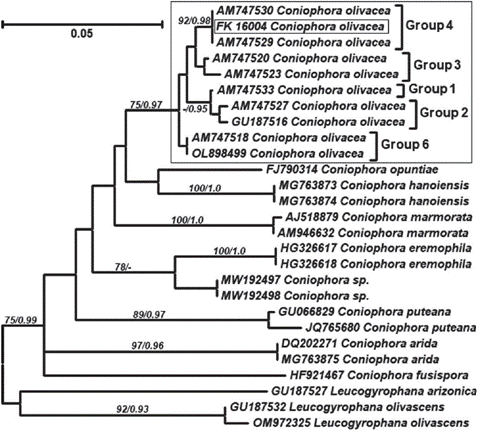
Fig. 1 Maximum likelihood tree showing significant bootstrap support (>75%) and significant Bayes- ian PP (>0.95). Intraspecific groups indicated in C. olivacea correspond to the cryptic clades defined by Kauserud et al. (2007). Group 5 was not recovered due to the lack of representative sequences.Fig. 1. Árbol de Máxima verosimilitud con valores significativos de bootstrap (>75%) y PP Bayes- ianaas (>0.95). Los grupos intraespecíficos indicados en C. olivacea corresponden a los clados crípticos definidos por Kauserud et al. (2007). El grupo 5 no está representado debido a la falta de secuencias observed as well as vessels with abundant hyphae.
Tangential sections made evident that rays are almost exclusively uniseriate. Vessel elements with oblique septa and opposite, scalariform intervessel pits, and with a large number of hyphae were also observed. These anatomical characteristics are coincident with Nothofagus dombeyi (Mirb.) Oerst wood, which has diffuse porosity, solitary and grouped vessels, in ra- dial, multiple, short and long series (Rivera, 1988). In tangential section, N. dombeyi presents oblique septa, and in radial section, simple perforation plates bordering each other, with subterminal, opposite (often scalariform) intervessel pits with circular to elliptical areolae. Rays are almost exclusively uniseriate, with procumbent cells (Tortorelli, 2009).
In transverse sections, the noninfected wood tested positive for lignin, where the fibers and rays are stained to a pink/red color (Fig. 2A), while in the wood col- onized by C. olivacea no positive reaction to lignin was detected. The fibers showed thin and deformed walls, and some showed cellular contents. The rays showed ocher colors with some reddish contents (Fig. 2B). In tangential longitudinal sections of noninfected wood, the fibers and walls of the vessels were stained to reddish/pink, evincing the presence of lignin (Fig. 2C). The wood showed a brownish coloration as well as no positive reaction for lignin (Fig. 2D).
Fibers of native (noninfected) tissues in the cross-section showed lignin pres- ence, as evinced by the blue color, indicative of this compound, whereas the rays showed a green coloration (a weaker reaction for the presence of lignin) with dark contents. The deposits in some vessels displayed a pink reaction indicating the presence of carbohydrates (Fig. 2E). In the tangential longitudinal section, the walls of the vessels showed intervascular pits stained with color blue as well as fibers, indicating the presence of lignin (Fig. 2G).
The cross-section of the infected wood showed fibers with thinner cell walls and rays with enlarged and ocher-colored contents that evince an advanced state of degradation. The vessels were observed to be translucent (white) due to the loss of carbohydrates and the absence of other contents (Fig. 2F). The situation was similar in the tangential section, where deformed vessels were also observed (Fig. 2H).
EMN of C. olivacea and potential distribution in Patagonia
The potential distribution model of C. olivacea exhibited a high predictive power, as shown by the AUC (μ: 0.902; SD: 0.114). The maps obtained from ENM showed how the habitat suitability is distributed globally (Fig. S3), including in the Pata- gonia region (Fig. 3). In general, suitable areas were located in Europe, North and South America, Australia, and East Asia. The model also predicted suitable areas in Argentina and Chile, which can represent geographic regions with an adequate combination of climatic and elevation conditions for the establishment of C. oliva- cea (Fig. 3). The areas with high suitability values (> 0.5), occupied approximately 629,053 km2 of the total area of Argentina and Chile (Fig. 3). According to this model, the main variables explaining the presence of the species were Bio12 (annu al precipitation) and Bio1 (annual mean temperature) contributing together with 81.6% (Table S3). Regarding the permutation importance, determined by random permutations of the values of each variable among the training points and back- ground data, Bio12 and Bio1 also had the maximum influence on the model and contributed 82.6% (Table S3). The result of the jackknife test showed Bio1 has the most useful information by itself and the most information that is not present in other variables (Table S4). In general, suitability was high in areas with an annual mean temperature (Bio1) between 0 °C and 10 °C, and precipitation values (Bio12) in the range of 500-1,500 mm3 (Fig. 4). The potential distribution extends in areas with low-temperature variation throughout the day (Bio2), and low-temperature change over the course of the year (Bio4) (Fig. S2); and in areas where the coldest temperatures are between -10 °C and 0 °C (Bio11) and low precipitation seasonality (Bio15) (Fig. S4). The most suitable areas are found below 1,000 m.a.s.l. with a mean annual temperature of around 10 °C (Fig. S5).
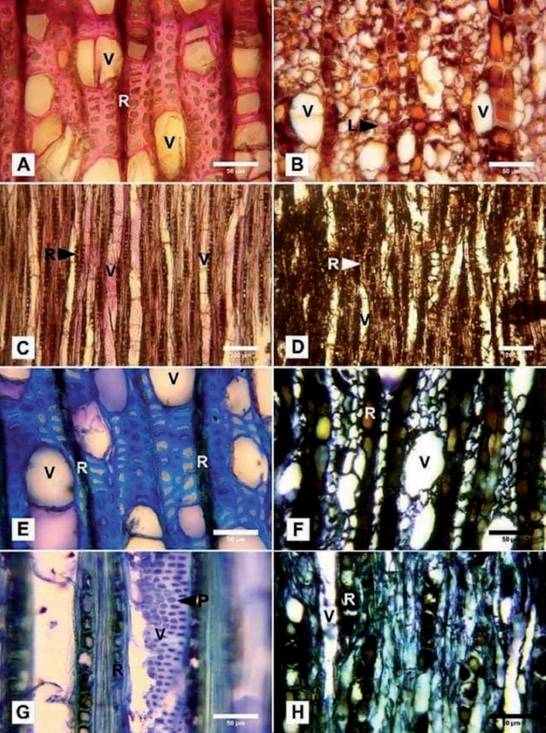
Fig. 2 Histochemistry of Nothofagus dombeyi uninfected wood (A, C, E, G) and infected by C. oli- vacea (B, D, F, H). A-D) stained with phloroglucinol. A) Crossection with fibers and rays showing a positive reaction (pink) to lignin. B) Thin, degraded walls without pink coloration (negative reaction to lignin) and rays with large cells and reddish-brown contents. Only a few fibers with lignin con- tents (L) are observed. C) Tangential longitudinal section of the vessel walls and fibers showing the presence of lignin. D) Negative reaction for lignin indicating the action of the wood decay fungi. E-H) Detail of the cross-section stained with toluidine blue, where: E: the fibers and rays with dark contents are observed along with some pink vessels indicating the presence of carbohydrates. F) Thin, deformed and dark-colored walls. Ocher rays with dark contents deformed (enlarged) cells and translucent vessels due to the degradation of carbohydrates. G) Tangential longitudinal section of non-infected wood where fibers and intervascular pits appear stained in blue, indicating the presence of lignin. The rays show a greenish coloration (a weaker reaction for lignin) with darker deposits. H) Infected wood with the dark fibers of thin walls and rays showing enlarged cells of dark cellular contents. (L) = lignin; (V) = vessels; (R) = rays; (P) = intervascular pits. Fig. 2. Histoquímica de madera no infectada de Nothofagus dombeyi (A, C, E, G) e infectada por C. olivacea (B, D, F, H) teñidas con floroglucinol. A) Sección transversal con fibras y radios mostrando reacción positiva (rosa) a lignina. B) Paredes delgadas, degradadas, sin coloración rosa (negativo para lignina) y radios de células grandes con contenidos castaño-rojizos. Solo se observan unas po- cas fibras con contenido de lignina (L). C) Sección longitudinal de las fibras y paredes de los vasos mostrando presencia de lignina. D) Reacción negativa a la lignina indicando la acción de hongos degradadores. E-H) Detalle de secciones transversales teñidas con azul de toluidina en las que, E: se observan fibras y radios con contenidos oscuros se observan junto con algunos vasos en rosa que indica presencia de carbohidratos. G) Sección longitudinal tangencial de madera no infectada donde las fibras y las punteaduras intervasculares aparecen teñidas de azul debido a la presencia de lignina. Los vasos muestran coloración verdosa (una reacción más débil a la lignina) con depósitos oscuros. H) Madera infectada con las fibras de paredes delgadas junto con radios que muestran células de contenido celular oscuro.
DISCUSSION
The highest habitat suitability (> 0.8) for C. olivacea seems be limited to humid forests in latitudes higher than 35° on the Chilean slope of the Andes. On the Ar- gentinean slope, areas of high suitability are more restricted, and limited to humid valleys in the provinces of Chubut, Santa Cruz and Tierra del Fuego. The rain sea- sonality seems to be decisive, slightly lowering the habitat suitability in the rest of the Andean Patagonia. This variable has shown to be also very influential, explain- ing the distribution of agaricoid fungal species in this region (Romano et al., 2017). However, the whole Andean Patagonia displays an also alarming suitability value above 0.6, in a region with widespread use of wood as main construction material and where wood production is one of the most important economic activities.
This high suitability supports the recognition of this record as an alien species. The chances of this species staying unnoticed despite the huge sampling efforts in the last decades are very low; especially considering the intensive campaigns looking for corticioid fungi in this region (Gallo et al., 2021). Also, the fact that the location of the collection studied has a long history of forestry and forest management speaks for a recent introduction.
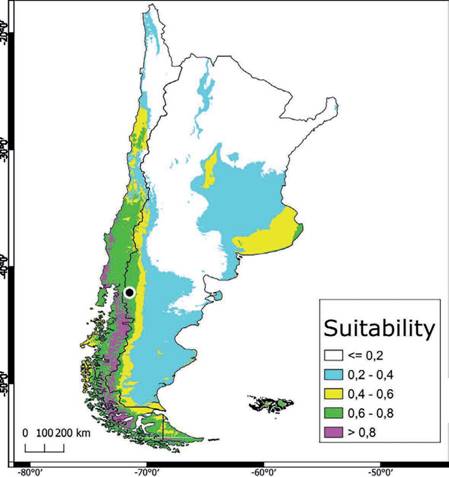
Fig. 3 . Distribution of suitable habitat of C. olivacea in Argentina and Chile. The suitability value represents the predicted distribution probability (in logistic value) for climatic and elevation layers used in the niche modeling. Specimen of C. olivacea collected in Patagonia marked with a black dotFig. 3. Distribución de hábitat idóneo para C. olivacea en Argentina y Chile. Los valores de idoneidad representan la distribución de probabilidad predicha (en valores logísticos) para las capas climáticas y de elevación usadas en el modelado de nicho. El especimen de C. olivacea colectado en Patagonia se marca con un punto negro.
An important subject to discuss here is the potential role of C. olivacea as a wood pathogen. There are some articles showing its role as a “butt rotter” in temperate and cold forests (Worrall & Nakasone, 2009) and also as a branch and stem rotter (Aref ’ev, 1991). Although it is too early to suggest that this species might act as a pathogen in Patagonia, the fact that the Patagonian collection is associated with a native tree species, together with the high habitat suitability shown, indicates that this possibility should be considered. Evidence of delignification was found in the substrate of the studied voucher despite the fact that C. olivacea is a brown rot spe- cies. This suggests that attack by C. olivaceae may be a secondary colonization on a previously decayed material. Assays in living trees under controlled conditions would be a potential way of assessing the latter. Furthermore, a more intense in- spection of living trees in the areas surrounding the original location would provide invaluable information on this matter. The verification of potential pathogenicity on native trees would require immediate actions restricting the spread of this decayer, given the wide range of wood species where it can grow (Da Costa et al., 1961).
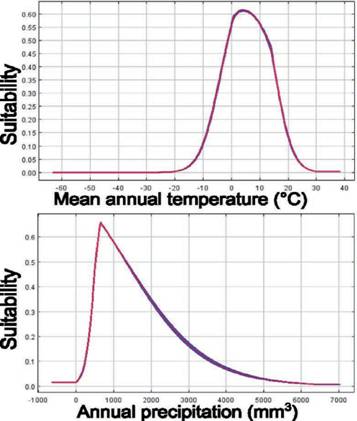
Fig. 4 Response curves of the most important environmental variables in modeling habitat distribu- tion for C. Olivacea. Mean annual temperature (Bio1) and Annual precipitation (Bio12). The suitability value represents the predicted distribution probability on a logistic scale. Fig. 4. Curvas de respuesta de las variables más importantes en el modelado de distribución de hábitat para C. olivacea. Temperatura anual media (Bio1) y precipitación anual (Bio12). El valor de idoneidad representa la distribución de probabilidad predicha en escala logística.
Finally, genotyping of more collections would allow analyses that take into ac- count the intraspecific diversity proposed by Kauserud et al. (2007). Such analyses are currently unviable given the number of registers needed to perform a supported prediction of the distributional range.














