INTRODUCTION
Bioinformatics is proving to be of great value for data manipulation in the field of biology. This area of knowledge encompasses all aspects related to the acquisition, processing, storage, distribution, analysis and interpretation of biological information. By unifying concepts and techniques from mathematics, statistics and computer science, it can lead to tools capable of extending the understanding of possible biological implications arising from genomic data [1].
Among the applications of bioinformatics, we find the sequencing and annotation of genomes, being the first responsible for providing the nucleotide composition of the genome of an organism. This technique is carried out by equipment from Illumina1, Ion Torrent2, Pacific Biosciences (PacBio)3 and Oxford Nanopore Technology (ONT)4, for example. This equipment allows the reading of text files containing DNA fragments called reads, which must be organised in order to represent the organism's genome. This procedure is called genome assembly, the most common approach being de novo, in which the genome is reconstructed exclusively from the overlay information of the reads. The sequences resulting from genome assembly are called contigs. For prokaryotic organisms, genome assembly can be performed by considering another genome as a reference model to guide the mapping of reads or by rearranging the contigs resulting from previous assembly [2,3]. Finally, the contigs are organised to form scaffolds, which will be used to perform genome annotation. Some examples of software used in this initial stage are: Velvet [4], SPAdes [5] and Trinity [6].
After sequencing, it is possible to obtain structural and functional information about the genome under investigation, as well as data that contribute to the evolutionary knowledge of organisms, innovations in diagnostic methods, new drugs, vaccines, possible means of prevention and more effective treatments against diseases or pests, and many other applications. Such information is obtained through annotation, which can be understood as a multilevel computational process involving nucleotides, proteins and processes [7].
In this context, for a project dedicated to genome assembly and annotation to develop successfully, it is necessary to initially define the application platform and the operating system to be used. After acquiring the equipment, translating the demands of a project into the specifications of a server (hardware) is fundamental to dimension the current needs. Depending on the chosen applications, it is important to carry out an analysis of the processing volume, memory usage and disk space consumption. For assembly, execution times and memory requirements increase with the amount of data. Therefore, there is a positive correlation between genome size and execution time and memory requirements [3]. For example, according to the company DNA STAR, which specialises in genomic sequencing, the organism Saccharomyces cerevisiae has a genome of approximately 15 MBases, thus requiring approximately 20 Gb of RAM, with 1 GB of RAM per MBase of genome length being recommended. These values can also be changed depending on the sequencer chosen and its mode of operation.
Genome assembly and annotation are purely computational procedures, usually performed by software without a graphical interface in a UNIX environment, which can be challenging for beginners in this environment. In addition, the results produced by one tool are not always in a format that can be used in the next tool in a workflow. In the literatura, it is possible to find articles that report the steps and procedures necessary for the execution of genomic assembly and annotation, and also discuss the peculiarities involved for each type of organism. In this regard, Zhou et al. [8] present the appropriate tools for genome assembly according to the sequencing approach. Keith [9] presents a compilation of methodological aspects related to genomic sequencing, assembly, annotation, data management, protein analysis and phylogenetic analysis. In the same vein, Ekblom and Wolf [10] analyse the workflow of a genomics project, from experimental procedures in the laboratory to the resulting applications of genome projects. In addition, Del Ángel et al. [3] point out important aspects that need to be analysed at the different stages. However, no article addresses the computational-technical difficulties related to the software execution environment.
In this context, many researchers in the field of life are discouraged to perform purely in silico procedures, either by technical-computational aspects or by the beginning of Bioinformatics as a science in the country. Although the installation of the software seems a simple task, during the assembly and annotation project of the fungus Penicillium equinulattum, carried out by researchers from the University of Caxias do Sul, some peculiarities were noticed that are not described in the installation tutorials and that, despite having relatively simple solutions, require significant time to diagnose. Thus, this article addresses technical aspects related to the Information Technology Infrastructure for its application in genomic assembly and annotation projects by researchers in the field of life sciences.
MATERIALS AND METHODS
The methodology consisted in the elaboration of an orientation workflow for the installation and configuration of a computer server dedicated to genomic assembly and annotation. In this regard, this section describes the software and hardware selection steps.
2.1 Genome Assembly and Annotation Tools
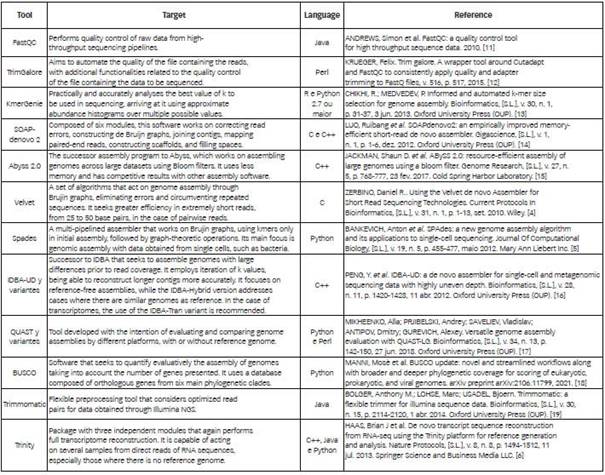
The choice of tools was based on the experience of the team involved in the P. echinulatum fungal assembly and annotation project, which includes researchers in the areas of Biology and Computer Science. In this sense, software or pipelines were sought to reduce computational difficulties, as shown in Table 1.
2.2 Hardware and operating system characteristics
For the development of the proposed configuration workflow, the hardware resources used were configured as follows:
Processor: Intel® Xeon® CPU x5650 at 2.67 GHz x 12;
Installed memory (RAM): 65.94 Gb;
System type: 64-bit system, X64-based processor.
Operating System: Ubuntu 20.04.2 LTS.
Among the operating systems, three stand out: MAC OS, Windows and Linux distributions, the latter being the most recommended for bioinformatics research purposes. Although less intuitive for users unaccustomed to open source systems, the variants present great flexibility and tools for practical purposes in the field. In addition, there are images and software developed for Linux that allow the automatic installation of programmes of interest or even help in the process, as is the case of Dugong and Unipro UGENE. There are many options for distribution, choosing Ubuntu in this article, because of the greater amount of information available on how it works and also because of the number of tools that support it.
2.3 Setting up the computing environment
The configuration of the computing environment involved the installation of the operating system, the configuration of the network environment, the firewall and the user account permissions. The chosen layout makes these steps clear and objective, without presenting major problems for the user. The greatest degree of difficulty is found in the use of the terminal, which is typical of Linux systems, but there is a wide range of manuals and guides on this in the virtual environment.
The next step must be the installation and configuration of the sequencing software, along with the necessary permissions to be able to operate fully. With these duly installed, it is possible to act on the management and processing of the memory dedicated to each one, finalising the configuration of the structure necessary to carry out the research related to the genome project. The software was chosen to cover the main steps of the sequencing process. FastQC [11], TrimGalore [12] and Kmergenie [13] were used for pre-processing. For assembly, SPAdes [5] and IDBA-UD [16] software were used to compare the difference between constant k-value and iterative k-value assemblies. Data evaluation was performed using Quast [17] and BUSCO [18] with reference genome. As for the annotation, only the initial part, referring to the structural annotation, was performed. The Augustus software [21], supported by the BLASTP tool [33], was used for this purpose. The datasets used are from the Staphylococcus aureus organism available on the SPAdes software homepage [5], due to the existence of previous results, forming a partial validation mechanism for the process.
RESULTS AND DISCUSSION
Initially, it is important to stress the great importance of creating a PATH variable to insert the programs to be installed and their dependencies, as well as the extremely important paths, such as the system's base folders. There are different ways to accomplish this task, such as using the "export" command in the terminal, which modifies the PATH variable only for the terminal in question, or modifying the file itself, in order to keep the changes. Regardless of the method chosen, there is a large source of detailed information on the process.
The installation of the mentioned software is similar. FastQC [11] and TrimGalore [12] are obtained from the same site. In both cases, the compiled .zip file is downloaded and extracted into the target directory. The program can be run via the terminal with the FastQC [11] folder directory defined or via PATH by the line “. /fastqc" or "fastqc", respectively. Attention should be paid to the Java version installed, because via PATH, other variants can be influenced so that the JRE has to be updated. It is best to install only the necessary dependencies according to a predefined list of programs to be used and their requirements. In the specific case of Java, you can run the command "java -version", obtaining the current version, and then "update-java-alternatives --list", which returns all the installed versions, inserted in the $PATH paths. Compared to the software version limiter, you can decide to keep it or change it using the line "sudo update-java-alternatives path/from/version".
TrimGalore [12] is executed by "trim_galore", either in the software's own path or via PATH, and its installation and execution did not present any other problems. In the case of Kmergenie [13] the download is via a "tar.gz" file. To perform the extraction, forward the folder to the destination directory and then enter the command "tar -xvf /directory", or similar. Then change the directory to the extracted folder and run the command make, which effectively installs the program, later executed via “. /kmergenie" or "kmergenie".
QUAST [17], despite having a browser version, can be installed, presenting in this format more variability and greater user control over the available parameters. Its installation is recommended, in order to enhance the learning of the tool and the understanding of the process as a whole. To work properly you need Python 2.5 and higher or Python 3.3 and higher, GCC 4.7 or higher, Perl 5.6.0 or higher, "GNU make" and "ar" and also the "zlib" package. It can be installed by the source file available on your site or by package managers, such as pip and linuxbrew. In the case of the source file, extract it into the target directory and then run the command line "quast.py" or "./quast.py" through the terminal. If it returns the software options, it is ready to use. To test the efficiency of the installation on a larger scale, run the command "quast.py -o /directory/to/results /quastdirectory/test_data/contigs_1.fasta". After checking the final message, which indicates the number of errors and warnings that appeared in the process. If no errors or warnings are returned, the program is ready for use. However, if it returns, you should check the "quast.log" file in the results directory to try to understand and resolve the problems. One warning that appeared several times during the development of this article was: "Cannot draw diagrams: python-matplotlib is missing or corrupted", referring to the lack of a specific python package, matplotlib, which is used in the creation of the graphics. After several tests of possible resolutions, a final and relatively simple conclusion was reached. The package in question was in a different version of Python run by QUAST [17]. To quickly solve the problem, the version of Python run by QUAST [17] presented in the initial lines of the terminal after executing the quast.py command was checked and found to belong to Python2. The command was then executed as "python3 quast.py", forcing it to run on Python3, the version on which the package was installed. This problem reinforces the importance of installing only the necessary dependencies when possible. You can try to work around this by assimilating the package to the Python2 version as well, but this would take more time and could cause obstacles in the future.
Another error encountered shows the output "'cgi' has no 'escape' attribute". To fix it, pay attention to the last line of the terminal before the error appears that interrupts the process and the directory it presents. Then find the file in the specified directory by opening it. Below the line "import cgi", add the line "import html", then find the line "cgi.escape", delete it and replace it with "html.escape". With these changes, QUAST [17] will run normally again. The latter problem seems to be more common when installing via pack manager pip. One interesting point worth mentioning is the fact that following this installation path for QUAST [17] also acquires the "--glimmer'' function, which already performs an initial prediction on the parsed sequence.
BUSCO [18] has several installation options, similar to those presented below for the annotation software and, as such, presents an extensive list of dependencies. It is executed by the busco command and has the configuration file call. This file contains information necessary for the proper functioning of the program, which must be edited before running it. It is recommended to specify all possible and relevant paths and parameters, thus ensuring a better performance of the program. An important point to note is the existence of optional and function-specific dependencies. In this case, it is up to the researcher to consider whether they are necessary or can be omitted in this project.
When it comes to annotation-related software, a more dense field is entered. For the most part, they require a large number of installed dependencies, which may include other annotation software, as in the case of Maker. As a counterpoint, they often feature web versions, which have limitations, but provide a more intuitive mechanism and multiple modes of installation. The routes involving package managers, pip and anaconda, include the addition of the necessary dependencies, making the process quicker and easier. Another route available is through the docker images, mostly executed by the command "compile docker" in the terminal directory containing the so-called Docker file. It should be noted that the above options often have limitations linked to specific functions, which are outlined in detail in the read-only files accompanying the software. The final alternative is to acquire the software via source code. All paths have advantages and disadvantages, and it is up to the user to gauge their needs and capabilities and, based on these, choose the right direction. It is recommended that you carefully read the websites and read.me files of the programs to be installed, as they contain an installation guide for different operating systems and information necessary for the correct execution of their code.
A good starting point is with Augustus and HMMER, as these, as well as individual programs that present interesting and extremely important results for structural annotation, are dependencies of other more "complex" programs such as BUSCO. In some programs, such as Augustus, you can change files, in this case "common.mk", setting COMPGENEPRED, Add SQLITE and Add MYSQL to "false", to obtain a simple and fast version of the program, which does not lose performance, only some areas of performance that may be dispensable for certain genomes.
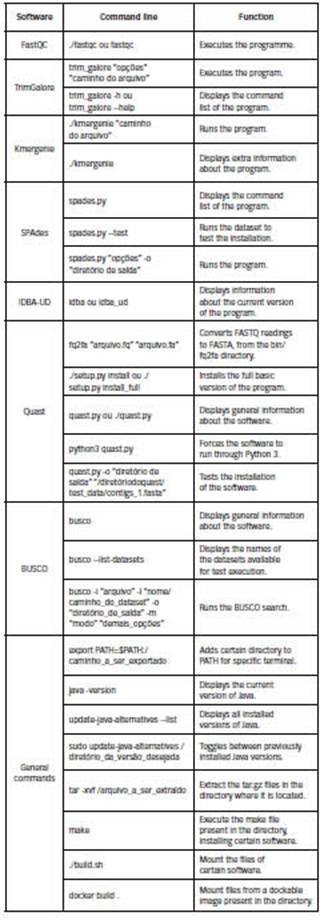
Due to the increase of dependencies and the diversity of applications, numerous version conflicts and errors can occur where already installed programs are missing because the directories are not included in the PATH variable. In programs with higher installation complexity, it is possible to perform a dependency check-list or even run the "make" command or the corresponding command that starts the assembly of the program, and wait for it to fail, because when this happens it issues an alert. .identify required files that are not specified. The executable files of the programs are installed in the /bin folder and, in case they are missing, you should consult the /src directory, always in the main folder of the software. So, in general, these are the paths to be added to the PATH variable. Below, you can see a table with the main commands of each chosen software, as well as some of general interest.
The part related to functional annotation will not be addressed in this article due to the high complexity present in this environment. To get started in this area, it is suggested to adapt to the use of BUSCO [18], due to the amount of dependencies and variable indexing system through the configuration file. In this way, the user will start to get used to the more computationally dense working environment and can then move on to more specific search and annotation programs. The use of the Maker programme can also be of great value to start the annotation process in general, as this software brings together several different steps in favour of the creation of genomic databases. However, its installation and execution is very complex. It works through four configuration files, as opposed to the single central file of Busco, and covers an extremely high number of dependencies, both mandatory and optional. In addition, it needs other programs, such as Apollo, to understand and study the results presented, programs that also have a more difficult installation flow.
CONCLUSIONS
The three general steps present very different degrees of complexity. Data pre-processing requires an understanding of the organism to be sequenced and how the experimental part of the process works. Basic level sequencing can be performed without much knowledge about the object of study, but optimal levels can be achieved with the addition of the same. In the case of annotation, it is essential to have extensive knowledge of the whole process logistics, both of the species and its phylogeny and of the programme to be implemented and the practical part of the study. This part of the project also often includes manual annotations, which requires special depth and focus.
At all levels of genetic sequencing, attention must be paid to the guides and text files provided by the programme developers, however, it is mainly the structural annotation programmes that require time to be set aside for reading. Through these you will discover the main lines of execution and the typical customisation options of the software, as well as its peculiarities. In addition, following or simply visiting the central Github page of the program, when available, can be extremely useful to identify possible bugs.
The comparison of assemblers, even if only between two programs, showed the essentiality of project planning, where software is chosen according to the needs presented. However, IDBA-UD [16] returned longer contigs, where the total length found for the genome was 2996254 base pairs, higher than that of SPAdes [5], which was 2977217 base pairs. In addition, the misassembled contigs presented by SPAdes [5] were more than five times longer than those presented by IDBA-UD [16]. Finally, the fraction of the genome constructed was 0.536 % higher through IDBA-UD [16], resulting in 98.799 %. However, it uses significantly more memory and time, and takes up to twice as long as SPAdes [5]. In view of the BUSCO analysis [18], the difference between the two assemblies was only 1 BUSCO, which appears fragmented in the assembly via SPAdes [5]. Therefore, between the two programs presented, SPAdes [5] is more economical and practical, while IDBA-UD [16] is denser and requires more powerful configurations, but is able to sequence with higher apparent accuracy.
On the structural annotation side, you could get interesting results with Augustus, although it runs at a basic level and without all the options that can be added to your pipeline. From the gene and protein sequences found, through BlastP [33], it was possible to generally evaluate the success of this portion. BlastP [33] identified different sequences presented by the programme as already studied sequences of the test organism, S. aureus.
The field of in silico gene sequencing is challenging for researchers who are not embedded in the environment. However, there is a growing number of tools that aim to facilitate the interaction of multiple user profiles from different projects. There is a great deal of flexibility and variability of programmes, allowing for an increasingly personalised choice appropriate to the resources available. By studying the theory and practice of the process and seeking to learn about the wide range of software available, satisfactory results can be achieved, which progress with the repetition and expansion of knowledge in the area.