Introduction
Chemical characteristics associated with the roughage used, such as dry matter content, concentration of water-soluble carbohydrates, buffer substances and populations of microorganisms present in the forage, and associated with the silage process bottlenecks, i.e. the silo filling time, compaction and sealing, are among the main factors that modulate the silage production process and modify the process and the characteristics of the silage 23,34.
The silage process is directly associated with the nature of the epiphytic microbial community, where bacterial diversity is the determinant factor of the fermentation pattern of silage. The microbial communities verified in forage crops before the silage process differ considerably in number and taxonomy from those quantified in silage 3,7,21.
Corn ( Zea mays L.) has adequate characteristics for good fermentation in a silo when harvested with the right dry matter content 16. However, due to the high concentration of water‐soluble carbohydrates (WSC), DM losses might occur during fermentation and when the silo is opened to withdraw the silage 16,26.
Thus, several corn silage inoculants have been researched in recent years, increasing knowledge about the dynamics of the action of bacteria in silage mass 31,32. The main additives were composed of microbial species of lactic acid bacteria (LAB), which are gram-positive, non-spore forming, strictly fermentative, anaerobic or aerobic tolerant, and acidophilic, and classified according to the type of hexose fermentation 16,18,28, which can be deferred. According to the species of Lactobacillus, Pediococcus, Leuconostoc, Enterococcus, Lactococcus and Streptococcus, classified as homolactic fermenters ( Lactobacillus plantarum) or heterolactic fermenters ( Lactobacillus buchneri), both producers of lactic acid and heterolactics can also produce acetic acid 17.
Lactobacillus buchneri has slow growth, with its effects observed from 45 to 60 days 20,29, which increases the aerobic stability of silage 8,24. The lactic acid and acetic acid produced are inhibitors of yeasts and moulds 20,31.
However, the formation of a large amount of acetic acid in silage does not present an advantage. This is related to the loss of energy and the reduction of the intensity of the pH drop in the silage 14,17. The further development of L. buchneri in silage mass may promote the increased aerobic stability of corn silage 30.
Through commercial products, the inoculation of silage occurs industrially from the dilution in water of the inoculant with strains of lyophilised lactic acid bacteria. The previous activation of the inoculant in reconstituted skim milk (RSM) prior to the ensiling process may increase the availability of active bacteria in the silage mass. The activation in RSM favours LAB in competition with other undesirable microorganisms and redirects the fermentation pattern and aerobic stability of the silage 27,31, promoting increased inoculation efficiency and higher yield and quality of the final product.
Knowledge of epiphytic microbial diversity can contribute to the understanding of the fermentation pattern of silage, as well as the phenomena arising from the exposure of this material to aerobiosis. Molecular biology techniques have been used to determine changes in the microbial community, favouring the intensity of responses about microbial diversity in silage and the effects of the inoculation of different strains of isolated LAB 13,26.
Accordingly, the aim was to evaluate the effect of inoculation with L. buchneri lyophilised or activated in RSM on LAB populations in corn silage.
Material and methods
Location and meteorological data
The experiment was conducted in the Forage Farming Sector of the Department of Animal Science of the Center of Agricultural Sciences, Federal University of Paraiba. The climate in the region is As’ (hot and humid), according to the Köppen classification. According to data from the Meteorological Station of the Agricultural Sciences Center of the Federal University of Paraiba, the average annual precipitation is 1400 mm; the average annual temperature is 24.5°C; and the average relative humidity is 80%.
Corn silage and treatments
The corn harvest was carried out at 97 days of age, when the grains were in the milky / pasty stage. The plants were harvested manually with an MS content of 26.2% and chopped in a stationary forage machine regulated to cut the forage into particles of approximately 2 cm and homogenised prior to inoculation and silage of the corn plant ( Table 1).
Table 1: Chemical composition and microbial populations of forage prior to inoculation and silage. Tabla 1: Composición química y poblaciones microbianas del forraje antes de la inoculación y ensilaje.
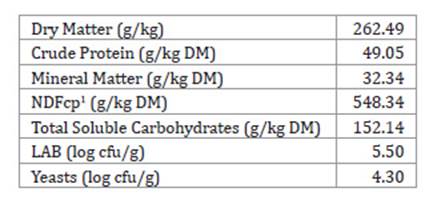
1 Neutral Detergent Fibre. corrected for ashes and proteins.
1 Fibra Detergente Neutra corregida para cenizas y proteínas.
Ensiling was carried out in tubular silos of PVC (15 cm diameter and 40 cm height), according to the treatments: silage without the inoculant ( in natura), silage with the lyophilised inoculant (SLI) and silage with the activated inoculant (SAI).
The inoculant with strains of L. buchneri (1 × 10 5cfu / g; Lactobacillus buchneri CNCM I-4323, Lallemand and Animal Nutrition) was applied according to the manufacturer’s recommendations (1 g / tonne fodder). The lyophilised inoculant was diluted in 100 ml distilled water and applied uniformly (2 ml / kg fodder) from a spray and constant mixture.
The inoculant was preactivated in 10% RSM 24 hours prior to ensiling. Skimmed milk powder (10 g) was solubilised in 100 ml of distilled water, and two grams of sucrose was added as an energy source for microbial growth, according to the methodology of Santos et al. (2008) . After growth, counts of lactic acid bacteria in the RSM were performed after 24 hours, as well as in the inoculant dissolved only in water, through the MRS culture medium 5 for Lactobacillus ssp. Populations of 2.1 × 10 8 and 4.5 × 10 6cfu / ml were grown in RSM growth medium and diluted in water, respectively. In 10 ml of RSM with reactivated L. buchneri, which was rediluted in 90 ml of distilled water, 2 ml / kg of natural material of this mixture was applied by spray.
The additives were mixed homogeneously to fill the experimental silos. Immediately, forage compaction was carried out in the silos, aiming to reach a specific mass of 600 kg / m 3 of natural matter. The silos were stored for 70 days at a mean temperature of 24°C before opening. Openings were performed at 1, 3, 7, 14 and 70 days after the silos were closed ( Table 2).
Table 2: Mean pH and concentrations of organic acids, ethanol and yeast of corn silage without inoculants and with lyophilised or activated microbial inoculants. Tabla 2: Valores medios de pH, concentraciones de ácidos orgánicos, etanol y levaduras de ensilajes de maíz sin inoculantes y con inoculante microbiano liofilizado o activado.

1 SLI = Corn silage with lyophilised microbial inoculant; 2 SAI = Corn silage with activated microbial inoculant; 3 Lat: Acet Ratio = Latic acid and Acetic acid ratio.
1 Ensilaje de maíz con inoculante microbiano liofilizado; 2 SAI = Ensilaje de maíz con inoculante microbiano activado; 3 Relación Lat: Acet = Relación de ácido láctico y ácido acético.
Quantification of lactic bacteria populations
The population count of lactic acid bacteria was performed according to the recommendations of González and Rodrigues (2003). Twenty-five grams of fresh silage samples were collected according to the defined opening periods, and 225 ml of sterile ringer solution was added and processed in a blender for approximately 1 minute. One millilitre of these mixtures was removed and pipetted to an appropriate dilution (10 -1 to 10 -9).
Plating was performed in duplicate for each culture medium. The populations were determined by the selective culture technique in anaerobic medium, where the culture medium MRS was used and incubated for 48 hours in an oven at 37°C, according to the methodology of De Man et al. (1960) .
The plaques considered susceptible to counting were those in which there were values between 30 and 300 cfu in a Petri dish. The plate means of the selected dilutions were then considered.
Lactic bacteria culture technique
After quantification of lactic acid bacteria populations using the pour plate technique in agar MRS culture 5, cultures were purified and cultured in a Falcon tube with 5 mL of MRS broth for 24 h at 37°C, and 10 cfu of each treatment was randomly selected 70 days after ensiling. The cultures were centrifuged for 10 minutes at 3600 rpm (rotation per minute) to obtain the cell pellet. The supernatant was removed, and 1 mL of saline (Ringer’s solution) was added and vortexed. Using a pipette, the entire volume of the pellet was transferred to the microtube and centrifuged at 6000 rpm for 3 minutes. The microtubes were stored in a freezer until DNA extraction was performed.
DNA extraction
DNA extraction from the isolates was performed using the Wizard Genomic DNA Purification Kit (Promega). The pellet was resuspended in 480 μl of 50 mM EDTA (Ethylic DiaminoTetracyclic Acid) in vortex. Fifty microlitres of lysozyme (50 mg / ml concentration) were added. The samples were incubated in a water bath at 37°C for 60 minutes and centrifuged for 2 minutes at 12,000 RPM, and the supernatant was removed.
After DNA extraction, DNA quantification procedures were carried out using Nanovue equipment (Nanodrop) at the Animal Biotechnology Laboratory of the Animal Science Department of the Federal University of Paraiba.
On average, the extraction resulted in a concentration of 1161.37 ng of DNA with 1.9 degrees of purity. After quantification, it was diluted to a concentration of 20 ng of DNA per μL.
Polymerase chain reaction (PCR)
Amplification of the 16S rDNA fragment of the isolates occurred with Primer 1492R (TAG G(C/T)A CCT TGT TAC GAC TT) and Primer p027F (GAG AGT TGA TCC TGG CTC AG) ( Heuer et al., 1997 ). The PCR reaction was performed in 0.2 mL tubes containing 50 μL of the reaction mixture: DNA (80 ng), 5X buffer solution (0.1 mol/L Tris-HCl, pH 8.0, 0.5 mol/L KCl), 1.5 mmol/L MgCl 2, pH 8.0; 0.2 mmol/L dNTP mix (Promega), Taq polymerase 2U (Promega), 0.12 μmol/L primer p027F 0.12 μmol/L, and 1429R 0.12 μmol/L (IDT Síntese Biotecnologia). The volume of the reaction mixture was filled to 50 μL with sterile ultrapure water. PCR was performed in a thermocycler 3Prime (Techne), and the reaction conditions employed in the PCR were: 94°C/5 min; 30 cycles (denaturation: 94°C for 30 seconds; 60°C for 30 seconds); polymerisation: 72°C/2 min; final extension: 72°C/5 min. An aliquot of 3 μl of the PCR product was mixed in 3 μl of the mixture: 1 μL Gel Red (Biotium) and 2 μL of pigment 6X Gel Loading (Promega) and analysed using agarose gel electrophoresis (1.2%) in buffer solution Tris-Borate-EDTA (TBE 1X). The gel was visualised under ultraviolet light, and images were captured using a gel system for photo documentation (MBS). The PCR product, a fragment of approximately 1500 bp, was sent to the Macrogen Company, Korea, for purification and sequencing.
Sequence analysis of the isolates
The sequences of the isolates were compared with those available in the GenBank database and aligned using the BLASTn algorithm (Basic Local Alignment Search Tool) ( http://www.ncbi.nlm.nih.gov/BLAST) for nucleotides. Sequences of the 16S rRNA gene that presented similarity equal to or greater than 95% were considered to belong to the same Operational Taxonomic Unit (OTU) 1.
Experimental design and statistical analysis
The experiment was carried out in a completely randomised design, with 3 treatments and 5 replicates per treatment, in each opening period (1, 3, 7, 14 and 70 days). The microbial counts were transformed into log10. Variance analysis and multiple comparisons of data were performed using the GLM procedures in SAS. The means were compared using the Kruskal-Wallis test.
Data on culture, DNA extraction, PCR and sequence analysis were performed and discussed through descriptive statistical analysis.
Results
Quantification of lactic acid bacteria in silage
Quantification of lactic acid bacteria populations was estimated at the different opening periods of the silos (1, 3, 7, 14 and 70 days). The lactic acid bacteria showed a rapid multiplication speed, reaching their highest development at 7 days of the fermentation process ( Figure 1, page 120).

SLI = Silage treated with the lyophilised inoculant; SAI = Silage treated with the activated inoculant. SLI = Ensilado tratado con inoculante liofilizado; SAI = Ensilaje tratado con inoculante activado.
Figure 1: Mean growth values of lactic acid bacteria in corn silage in natura and inoculated with Lactobacillus buchneri during different fermentation periods. Figura 1: Valores medios de crecimiento de bacterias lácticas en ensilaje de maíz in natura e inoculado con Lactobacillus buchneri durante diferentes períodos de fermentación.
Identification of lactic acid bacteria populations in silage
It was observed in this study that a greater number of the isolates (93.34%) were formed by lactic acid-producing strains of the species L. plantarum ( Table 3, page 120).
Table 3: Molecular identification of samples of bacteria of the genus Lactobacillus isolated 70 days after ensilage. Tabla 3: Identificación molecular de muestras de bacterias del género Lactobacillus aisladas a los 70 días del ensilaje.

With the sequencing of the 16S rDNA fragment produced by PCR using Primers 1492R and p027F, the isolates were closely related to L. plantarum. Figure 2, shows that the PCR products were visualised using agarose gel electrophoresis.
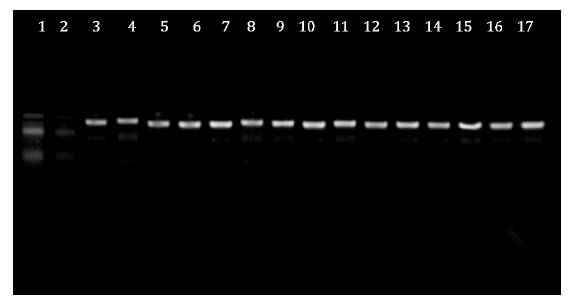
Figure 2: PCR amplification products for Lactobacilli. Line 1, Molecular marker (100 bp DNA ladder); Line 2, White; Lines 3 to 17, Lactobacillus plantarum. Figura 2: Productos de amplificación por PCR para Lactobacilli. Línea 1. Marcador molecular (escalera de DNA de 100 pb); Línea 2. Blanco; Líneas 3 a 17: Lactobacillus plantarum.
Discussion
The silage inoculated with preactivated L. buchneri had the highest values of lactic bacteria populations until the 14 th day of the fermentation period, reaching 11 log CFU / g in the forage at 7 days of fermentation ( Figure 1, page 120). These results corroborate those observed by de Santos et al. (2008) since there was the addition of a larger number of active bacteria, thus facilitating their multiplication. An indication of the presence of heterofermentative bacteria was the reduction of lactic acid concentration, the increase in acetic acid concentration and the reduction of the proportion of lactic and acetic acids. Consequently, the ethanol levels were low at the end of the 70 days of silage 6.
However, the addition of a group of microorganisms as additives at the end of the silage process may not be present. These microorganisms promote changes in the pH value, and the presence of several organic compounds, such as acetic, lactic, butyric, propionic, ethanol, CO 2, and antimicrobial and/or bacteriostatic compounds, also affect the interaction between the various groups of microorganisms 14.
These modifications were desirable considering that the pre-silage material had a dry matter content (26.2%) below that recommended for good-quality silage 16. According to Danner et al. (2003) , lactic acid formation is fundamental to reducing the pH of the silage, causing the selection of microorganisms in the silage mass. However, acetic acid also needs to be formed to ensure that the silage mass has an adequate pH of 3.8-4.2 16, thus providing control of moulds and yeasts. However, when the pH value is reduced too much, it may favour the development of these microorganisms, which causes a decrease in aerobic stability 20.
Although there was no reduction in yeast in the corn silage with lyophilised or activated microbial inoculant, the presence of acetic acid has antifungal action, which reduces the production of other compounds, such as ethanol 22,36, as observed in this study. According to Sadiq et al. (2019) , there is a consensus from the scientific community that organic acids are the main antifungal metabolites of LAB.
Organic acids in their dissociated or undissociated form are lipophilic in nature and thus readily diffuse across the fungal cell membrane and accumulate in the cytoplasm, reducing activity or leading to yeast death. However, acetic acid has higher inhibitory activity against fungal growth compared to lactic acid. Although the LAB populations of the silages treated with the lyophilised inoculant showed lower growth compared with the activated inoculation, they stabilised at 70 days with populations above 9.0 log cfu/g forage. The values were similar in silage with the inoculant and higher than in silage without the inoculants ( Figure 1, page 120). This difference may be due to a peak in LAB development at 7 days of silage, when there may have been intensified substrate competition. Probably, the activation of the lyophilised LAB with RSM caused this difference in the population on the 7 th day of silage, which remained until the 70 th.
The fermentative process of silage is complex and involves many species of LAB and their interactions. The use of specific inoculants is indicated to dominate or overcome the number of epiphytic lactic bacteria present in the forage, either to improve the fermentation process 19,32 or to increase the aerobic stability of silage 37. However, the increase in LAB in the ensiled mass promotes the greater availability of specific microorganisms. The greater or lesser degree of development of these bacteria depends on the conditions of the medium 9.
Inoculation of the silage with activated heterofermentative LAB ( L. buchneri), facultative aerobes, produces lactic acid and acetic acid. These characteristics cause a rapid proliferation of the same in the ensiled mass and cause an adequate reduction of the silage pH, controlling the growth of yeasts and moulds ( Table 2, page 118). Thus, the ensiled mass can provide greater development of epiphytic bacteria, such as L. plantarum, which is homofermentative 15,28.
The pH of the ensiled mass rapidly decreased due to the homofermentative LAB multiplying rapidly, producing more lactic acid. Lactobacillus buchneri bacteria limit the growth and metabolism of lactic acid degradation as a strategy to maintain cell viability ( Table 2, page 118) 20,29. The activity of L. buchneri was evidenced by higher acetic acid production. In the 70-day fermentation period, the selected isolates of inoculated corn silage were not identified ( Table 3, page 120). Possibly due to the greater fermentative capacity in the corn silages, L. plantarum dominated the fermentation of the silage. Guo et al. (2018) observed similar behaviour in alfalfa silage, including L. buchneri, which is tolerant to acidic environments and uncompetitive compared to other LAB species.
The high frequency of L. plantarum in corn silage promoted the effective fermentation of lactic acid, rapidly reducing the pH of the ensiled mass and preventing the development of microorganisms deleterious to silage, such as Clostridia and enterobacteria. In addition to decreasing the pH from lactic acid production, L. plantarum can also inhibit the growth of filamentous moulds through the production of antifungal activity 2. However, this characteristic is more likely for heterofermentative LAB species 25.
In this study, strains of Lactobacillus paraplantarum (6.6% of isolates) were also identified in corn silage inoculated with RSM pre-activated bacteria ( Table 3, page 120). Zhang et al. (2017) observed the presence and predominance of L. paraplantarum in LAB populations in corn silage with high humidity.
Wang et al. (2017) , characterising isolated lactic acid bacteria and their effects on the fermentation of silage, verified the growth of L. paraplantarum limited to 10°C and pH 3.0. According to the authors, the growth of L. paraplantarum strains in low pH and temperature environments confirmed the resistant and acidic nature of this species. At the end of the 70-day silage period, the dynamics of the microbial population provided the prevalence of the largest population of L. lantarum among LAB. These findings corroborate the findings of Guo et al. (2018) , who reported that alfalfa silage inoculated with L. buchneri changed the population profile of the epiphytic LAB species on the 60th day after silage. The same authors stated that the L. plantarum population predominated by more than 90%, but they failed to confirm the cause of this change with certainty and suggested more studies related to the metabolomics of LAB.