Microsporidia are obligate intracellular parasites with a remarkable ability to exploit a wide range of inverte- brate and vertebrate animals, causing from cryptic, benign infections to massive infections with extensive damage leading even to host death5. These organisms were previ- ously considered ''primitive’’ protozoa; however, molecular phylogenetic studies as well as biological evidence have demonstrated that microsporidia are related to the kingdom Fungi, either as a basal branch or sister group5. Although there are 8 genera and 14 species able to infect humans, two species are the most commonly identified, Enterocy- tozoon bieneusi and Encephalitozoon intestinalis. Human infections are thought to be mainly zoonotic or waterborne and microsporidia spores are released into the environment with feces, urine, and respiratory secretions5. Immuno- competent individuals have asymptomatic or self-limited infections, occasionally reported as ''traveler’s diarrhea’’; however, AIDS patients or other immunosuppressed individu- als suffer intestinal microsporidiosis with chronic diarrhea, malabsorption syndrome or severe disseminated disease5. The prevalence of this emerging opportunistic infection in human ranges from 1 to over 50%, depending on the geographic region, the methods applied for diagnosis and demographic characteristics of the studied population11. Albendazole is effective against many species including Encephalitozoon spp., but it shows low efficacy in control- ling E. bieneusi disease5.
Transmission electron microscopy (TEM) is regarded as the golden standard for establishing the diagnosis but is expensive, time-consuming and has low sensitivity. Light or immunofluorescence microscopy and molecular amplifi caron techniques, primarily using PCR, are now routinely used for diagnosis. As a matter of fact, PCR assays are sig- nificantly more sensitive than light microscopy and provide species differentiation, thus having an impact on treatment decisions3.
In Argentina, 120000 people live with Human immunod- eficiency virus (HIV)/acquired immune deficiency síndrome (AIDS) and 6500 new cases are reported annually. In turn, 27.5% of these people are diagnosed late; there- fore, they are often at risk of acquiring an opportunistic infection7. Nevertheless, there are fewepidemiological data on microsporidiosis13, and a reliable diagnosis is particularly necessary for AIDS patients with chronic diarrhea12. In this study, we standardized microscopy techniques as well as a nested PCR to detect microsporidia in stool samples2,8 so as to be further applied in diagnosis and epidemiological studies in public health in Argentina.
Stool samples (Sample A and B, non-HIV) from Rawson Hospital (Córdoba, Argentina) stock collection samples were used as negative controls and the E. bieneusi positive sample was provided by the Parasitology Laboratory of Hospital Gar- raham, Buenos Aires, Argentina. The research protocol was approved by the Ethical Committee for Research of Rawson Hospital, Córdoba, Argentina (number 05072018).
Chromotrope 2R technique was performed by following the method described by Weber et al.14, with slight modifi- cations. Briefly, stool samples were fixed with ethanol 70% (1/3 ratio) and then concentrated by the modified ethyl- acetate stool-concentration method4. Smears were fixed with methanol for 5 min and incubated overnight with chro- motope 2R stain, prepared as described by Weber et al. 14 After staining, slides were rinsed and de-colored by dipping in acid alcohol for 10s (moving the slide up and down) and stopped with 95% alcohol. Smears were successively dehy- drated in 95% alcohol (5 min), 100% alcohol (10 min) and xylene (10 min). In parallel smears, calcofluor white staining was also performed as previously described6.

Figure 1: Weber’s modified trichome (A and B) and calcofluor (C) staining showing Enterocytozoon bieneusi spores in feces. (A) Modified trichrome staining observed by light microscopy showing ovoid shaped-spores with a pinkish-red stained wall (arrows). Some spores show a pinkish belt-like stripe. (B) Calcofluor staining observed by fluorescence microscopy showing positive E. bieneusi spores (arrows) and a yeast (asterisk). Magnification 1000x. Bars = 5 pm.
To confirm microsporidia detection by light microscopy, we further performed a nested technique as previously described1 and modified by Santin-Duran et al.2,9,10. DNA extraction was performed by using the DNAeasy Blood & Tissue Kit (Qiagen, USA), according to the manu- facturer’s instructions. PCR was performed using a set of nested primers specific for E. bieneusi that ampli- fied the ITS and portions of the flanking large and small subunits of the rDNA (~400bp). The outer primers were EBITS3 (5'-GGTCATAGGGATGAAGAG-3') and EBITS4 (5'-TTCGAGTTCTTTCGCGCTC-3'), and the inner primers were EBITS1 (5'-GCTCTGAATATCTATGGCT-3') and EBITS2 (5'-ATCGCCGACGGATCCAAGTG-3')2. The reaction mixture (20^l) contained 2 ^l of buffer 10x (16mM MgCl2, 500mM KCl, 100 mM Tris-HCl; Dongsheng Biotech, China); 0.16 ^l of MgCl2 50 mM; 0.4 ^l of 10 mM dNTPs; 20 pmol of each primer;
2 ^l of 5 U/^l of Taq polymerase (Dongsheng Biotech); and 5 ^l of purified DNA. After denaturation at 94 °C for 3 min, the PCR samples were subjected to 35 cycles of amplifica ron (denaturation at 94 °C for 30 s, annealing at 57 °C for 30s, and elongation at 72°C for 40s), followed by a final extension at 72 °C for 10 min. Nested PCR conditions were
identical to those of the first run except that only 30 cycles were carried out with an annealing temperature of 55 °C. Pure and diluted (1/10,1/100 and 1/100) PCR products were used in the nested PCR. A negative control without DNA was included in all PCR sets. Furthermore, to rule out PCR inhi- bition, DNA amplification from Candida albicans was also performed in all samples (Supplementary Fig.). PCR prod ucts were subjected to electrophoresis in 2% agarose gel and visualized by staining the gel with ethidium bromide.
To determine E. bieneusi identity, the amplification product from the nested PCR was then subjected to a direct nucleotide sequencing reaction in both directions by using the inner-nested PCR primers (EBITS1 and EBITS2) in the Applied Biosystems® 3500xL Genetic Analyzer at Laboratorio Central de la Provincia de Córdoba, Argentina. Sequences obtained were aligned and inspected by using the MEGA software (version 6), and then compared with refe- rence sequences from the GenBank database by the BLAST analysis10.
Microsporidia are still difficult to diagnose even though significant progress has been made over the last few years. Nowadays microscopic or PCR-based detection techniques, which must be standardized in each laboratory4, are useful for diagnosis and genotyping. In this study, we standardized the most commonly used staining technique for the exami- nation of stool specimens: chromotrope 2R-based (Weber’s modified trichrome) stain14 and chemifluorescent optical brightening agent calcofluor white (that detects chitin in the spore cell wall)4. Figure 1 A shows pinkish red-stained microsporidia spores, with some of them showing a distinct pinkish belt-like stripe (1000x). The spores show a characteristic staining pattern and can be readily distin- guished from bacteria and fecal debris4,14. In addition, as described by Weber et al.14, calcofluor stained not only microsporidia spores but also yeasts and some feces debris (Figure 1B). In this regard, previous studies comparing the chromotrope staining technique with methods using chemi- fluorescent optical brighteners have shown that these tests are robust for routine use4. However, none of these micro- scopic methods are sufficient to determine microsporidia species; therefore, we also performed a nested PCR that is currently used to identify E. bieneusi genotypes and other microsporidia from stool samples. Figure 2 shows that nested PCR produced fragments of approximately 390 bp
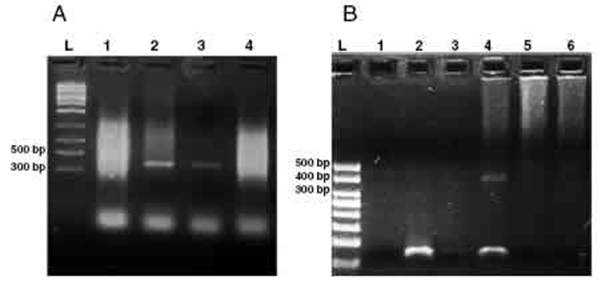
Figure 2: Agarose gel electrophoresis (2%) showing amplification products from the nested from ethanol-fixed stool samples. (A) PCR from Enterocytozoon bieneusi positive stool sample. L: molecular weight ladder. Lanes 1 -3: products of nested PCR (with inner primers ITS1 and ITS2) using diluted products from PCR as DNA template (Lane 1: 1/10, Lane 2: 1/100, Lane 3: 1/1000). Lane 4: PCR product using undiluted DNA template from PCR. (B) Nested PCR from different stool samples. L: molecular weight ladder, Lane 1: negative control (H2O), Lane2: negative stool sample A (PCR product diluted 1/100), Lane 3: negative stool sample B (1/100), Lane 4: positive stool sample (1/100), Lane 5: negative stool sample A (diluted 1/10), negative stool sample B (diluted 1/10).
(Fig. 2A and B). Furthermore, the best performance was obtained when products from the first amplification reac- tion (435bp, not shown) were diluted (1/100and 1/1000) to be used in the nested PCR (Fig. 2A).
On the other hand, this PCR technique amplifies the inter nal transcribed spacer (ITS) region of the rRNA, which has been useful in many studies for the identification of E. bieneusi genotypes1Accordingly, the sequence anal- ysis of PCR-amplified products showed 99.68% (319 pb) homology with genotype B of E. bieneusi (GenBankaccession number AF101198.1)10.
In conclusion, in this work we successfully detected E. bieneusi microsporidia from a human stool sample by using the Chromotrope 2R microscopic technique and a previ- ously reported nested PCR, which can also be applied for E. bieneusi genotyping 2,8-10 . Further studies are currently underway to define whether these techniques could be use ful for human microsporidiosis surveillance in Argentina.
Funding
This work was supported by the Secretary of Science- and Technology of the National University of Cordoba (Sec-retaría de Ciencia y Tecnología de la Universidad Nacionalde Córdoba, Argentina, SECyT-UNC, PROYECTO CONSOLIDARNo33620180100408) and the National Agency for Scien-tific and Technological Promotion - Fund for Scientific andTechnological Research (Agencia Nacional de PromociónCientífica y Tecnológica, Fondo para la Investi gación Cien-tífica y Tecnológica, Argentina, FONCyT, PICT 2015-1425).CJM is a fellow from SECyT-UNC (Grants for Socio-productiveTechnological Innovation - Becas de Inno vación TecnológicaSocio-Productiva, BITS 2018) and LSC is a member of thescientific career of the National Council for Scientific andTechnological Research (Consejo Nacional de InvestigacionesCientíficas y Tecnológicas, CONICET), Argentina.
Conflict of interest
The authors declare that they have no conflicts of interest.
Acknowledgment
We wish to thank Dr. Mónica Santin-Duran (United States Department of Agriculture, Maryland, USA) for providing PCR protocols and Dr. Sebastian Dambolena (IMVIB-CONICET, Cór doba, Argentina) for technical support.
Appendix A. Supplementary data
Supplementary data associated with this article can be found, in the online version, at












 uBio
uBio 

